12.4 – ANOVA from “sufficient statistics”
Introduction
By now you should be able to run a one-way ANOVA using R (and R Commander) with ease. As a reminder, You should also be aware that, if you need to, you could use spreadsheet software like Microsoft Excel or LibreOffice Calc to run one way ANOVA on a small data set. Still, there are times when you may need to run a one-way ANOVA on a small data set, and doing so by hand calculator may be just as convenient. What are your available options?
Following the formulas I have given would be one way to calculate ANOVA by hand, but it would be tedious and subject to error. Instead of working with the standard formulas, calculator shortcuts can be derived with a little algebra and this is where I want to draw your attention now. This technique will come in handy in lab classes or other scenarios where you collect some data among a set number of groups and calculate means and standard deviations. The purpose of this posting is to show you how to obtain the necessary statistics to calculate a one-way ANOVA from the available descriptive statistics: means, standard deviations, and sample sizes. In other words, these are the sufficient statistics for one-way ANOVA.
Note 1: In Chapter 11.5, we introduced use of summary statistics, ie, “sufficient statistics,” to calculate the independent sample t-test.
As you recall, a one-way ANOVA yields a single F test of the null hypothesis that all group means are equal. To calculate the F test, you need
- Mean Square Between Groups,
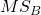
- Mean Squares Within Groups or Error,
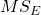
F is then calculated as
![]()
with degrees of freedom k – 1 for the numerator and N – 1 for the denominator. ![]() can also appear as
can also appear as ![]()
We can calculate ![]() as
as
![]()
where k is the number of i th groups, ni is the sample size of the ith group, ![]() refers to the overall mean for all of the
refers to the overall mean for all of the ![]() sample means.
sample means.
Next, for the Error Mean Square, ![]() , all we need is the average of the sample variances (the square of the sample standard deviation, s).
, all we need is the average of the sample variances (the square of the sample standard deviation, s).
![]()
ANOVA from sufficient statistics
Consider an example data set (Table 1) for which only summary statistics are available (mean and standard deviation, sd). The data set is for life span(days) for several inbred strains of laboratory mice from a study (. Sample size for each group was seven mice.
Table 1. Descriptive statistics female life span (days) for ten different inbred strains of mice (Mus domesticus).
| Strains | n | mean | sd |
|---|---|---|---|
| 129S1/SvImJ | 32 | 787.4 | 159.16 |
| A/J | 32 | 630.7 | 130.20 |
| BALB/cByJ | 32 | 734.4 | 154.43 |
| BUB/BnJ | 24 | 611.3 | 218.34 |
| C3H/HeJ | 29 | 724.1 | 131.48 |
| C57BL/6J | 29 | 855.7 | 185.34 |
| CBA/J | 30 | 622.9 | 181.95 |
| FVB/NJ | 26 | 750.3 | 230.11 |
| P/J | 32 | 676.0 | 178.82 |
| SWR/J | 31 | 831.9 | 181.31 |
Data from Yuan et al (2021; https://phenome.jax.org).
Spreadsheet calculations.
You have several options at this point, ranging from using your calculator and the formulas above (don’t forget to square the standard deviation to get the variances!), or you could use Microsoft Excel or LibreOffice Calc and enter the necessary formulas by hand (Table 2). You’ll also find many online calculators for one-way ANOVA by sufficient statistics (eg, https://www.danielsoper.com/statcalc/calculator.aspx?id=43).
Table 2. Spreadsheet with formulas for calculating one-way ANOVA from means and standard deviations from statistics presented in Table 1.
| A | B | C | D | E | F | G | H | I | |
| 1 | Strain | n | Mean | sd | squared | variance | grand mean |
=AVERAGE(C:C) | |
| 2 | 129S1/SvImJ | 32 | 787.4 | 159.16 | =B2*(C2-$I$1)^2 | =D2^2 | dfB | =COUNT(B:B)-1 | |
| 3 | A/J | 32 | 630.7 | 130.2 | =B2*(C3-$I$1)^2 | =D3^2 | dfE | =SUM(B:B)-I2 | |
| 4 | BALB/cByJ | 32 | 734.4 | 154.43 | =B2*(C4-$I$1)^2 | =D4^2 | Msb | =SUM(E:E)/(COUNT(E:E)-1) | |
| 5 | BUB/BnJ | 24 | 611.3 | 218.34 | =B2*(C5-$I$1)^2 | =D5^2 | Mse | =SUM(F:F)/COUNT(F:F) | |
| 6 | C3H/HeJ | 29 | 724.1 | 131.48 | =B2*(C6-$I$1)^2 | =D6^2 | F | =I4/I5 | |
| 7 | C57BL/6J | 29 | 855.7 | 185.34 | =B2*(C7-$I$1)^2 | =D7^2 | P-value | =FDIST(I6,I2,I3) | |
| 8 | CBA/J | 30 | 622.9 | 181.95 | =B2*(C8-$I$1)^2 | =D8^2 | |||
| 9 | FVB/NJ | 26 | 750.3 | 230.11 | =B2*(C9-$I$1)^2 | =D9^2 | |||
| 10 | P/J | 32 | 676.0 | 178.82 | =B2*(C10-$I$1)^2 | =D10^2 | |||
| 11 | SWR/J | 31 | 831.9 | 181.31 | =B2*(C11-$I$1)^2 | =D11^2 |
For this example, you should get the following
MSB = 219899.5 MSE = 31634.1 F = 6.951 P-value = 4.45E-09
Note 2: The number of figures reported for the P-value implies a precision that the data simply do not support. For a report, recommend writing the P-value < 0.001. But note — truncating the p-value diminishes the value to others who may wish to conduct a meta analysis using your work (Chapter 20.15)!
But, R can do it better.
Here’s how. Install the HH package (or RcmdrPlugin.HH for use in Rcmdr) and call the ?aovSufficient function.
Step 1. Install the HH package from a CRAN mirror, e.g., cloud.r-project.org, in the usual way.
chooseCRANmirror()
install.packages("HH")
library(HH)
Step 2. Enter the data. Do this in the usual way (eg, from a text file), or enter directly using the read.table command as follows.
MouseData <- read.table(header=TRUE, sep = ",", text= " Strain, n.size, average, stdev 129S1, 32, 787.4, 159.16 A.J, 32, 630.7, 130.20 BALB.c, 32, 734.4, 154.43 BUB.BnJ, 24, 611.3, 218.34 C3H.HeJ, 29, 724.1, 131.48 C57BL.6J, 29, 855.7, 185.34 CBA.J, 30, 622.9, 181.95 FVB.NJ, 26, 750.3, 230.11 P.J, 32, 676.0, 178.82 SWR.J, 31, 831.9, 181.31") #Check import head(MouseData)
End of R input
I know, a little hard to read, but from the MouseData to the end bracket ") before the comment line #Check import, that’s all one command.
Of course, you could copy the data and import the data from your computer’s clipboard in Rcmdr: Data → Import data → from text file, clipboard, or URL…
Note 3: Hint: for field separator, try comma; if that fails, try Tabs
Once the data set is loaded, proceed to Step 3.
Step 3. In our example, sample size was included for each group. Skip to step 4. If, however, the table lacked the sample size information, you can always add a new variable. For example, if we needed to add sample size to the data frame, we would use the repeat element function, rep().
MouseData$n <- rep(7, 10)
If you check the View data set button in Rcmdr, you will see that the command in Step 3 has added a new variable “n” for each of the eleven rows. The function rep() stands for “replicate elements in vectors” and what it did here was enter a value of 7 for each of the ten rows in the data set. Again, this step is not necessary for this example because sample size is already part of the data frame. Proceed to step 4.
Step 4. Run the one way ANOVA using the sufficient statistics and the HH function aovSufficient
MouseData.aov <- aovSufficient(average ~ Strain, data=MouseData)
Step 5. Get the ANOVA table
summary(MouseData.aov)
Here’s the R output
Df Sum Sq Mean Sq F value Pr(>F) Strain 9 1977732 219748 7.123 2.54e-09 *** Residuals 287 8853743 30849 --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
End R output
To explore other features of the package, type ?aovSufficient at the R prompt (like all R functions, extensive help is generally available for each function in a package).
Meta-analysis and sufficient statistics.
Meta-analysis (see Chapter 20.15 — Meta-analysis), can be performed using sufficient statistics from different tests. Combining sufficient statistics allows for a more precise overall estimate by effectively merging the information from multiple studies, which can be especially useful when individual studies have small sample sizes or lack a unified outcome measure. Common methods include creating summary effect sizes like the standardized mean difference or using p-value combination methods, while ensuring the underlying statistical assumptions are met through goodness-of-fit tests.
Fisher’s method combines the p-values from multiple independent studies into a single overall p-value to test a joint null hypothesis. The method’s test statistic is calculated as the sum of the negative log-transformed p-values. P-values from ![]() number of tests are combined; each p-value,
number of tests are combined; each p-value, ![]() , contributes to a variable,
, contributes to a variable, ![]() , with
, with ![]() degrees of freedom. The chi-square distribution is used to evaluate the significance of the Fisher’s method test statistic.
degrees of freedom. The chi-square distribution is used to evaluate the significance of the Fisher’s method test statistic.
Fisher’s method is available in several R packages (eg,poolr package); here, we’ll just create a simple R code.
Note 4: Not to be confused with Fisher’s exact test, Chapter 9.5.
Limitations: Methods that only use p-values or test statistics may not adequately address the effects of heterogeneity and potential publication bias among studies. Because many older studies do not report effect sizes and confidence intervals, using test statistics or p-values is often the only available method to combine their results.
Example.
One hypothesis about longevity is that the greater the genetic diversity can lead to longer life. The mice in Table 1 were all from inbred mouse strains. By definition, genetic diversity is restricted in inbred mice, compared to outbred strains. Jackson Labs produced a “Diverse Outbred” line of mice, and a few individuals have lived nearly five years, well past the typical 1 – 2 year lifespan of mice (Cohen 2017, Mullis et al 2025). Thus, a natural comparison is to ask: is there a difference in average lifespan between the DO and inbred mouse strains? We’ll run a series of one-sample t-tests against the mean lifespan for the diverse outbred stock of mice, save the p-values, and apply Fisher’s method. While not strictly a meta-analysis, which would draw from multiple studies, this example outlines the basic steps to take given a series of hypothesis of the same kind.
Note 4: Used another R package, BSDA, for a series of one-sample t-tests. DO mean of 794.5 + 262.85 days (n = 192 females), calculated from supplement data in Mullis et al 2025). R code example:
do_avg= 794.5 # dataset MouseData tsum.test(average[1], stdev[1], n.size[1], mu=do_avg)
Abbreviated R results
t = -0.25235, df = 31, p-value = 0.8024
Repeat for all ten strains from Table 1; p-values were saved to Table 2.
Obviously, a for loop would be appropriate here! The R code extracts p-values from each of the ten tests and saves to p_values object.
p_values <- numeric(length(average))
for (i in 1:length(average)) {
suppressWarnings({
test_result <- tsum.test(
mean.x = average[i],
s.x = stdev[i],
n.x = n.size[i],
mu = do_avg
)
p_values[i] <- test_result$p.value
})
}
print(p_values)
I’ve wrapped the tsum.test in a suppressWarnings({}) line to prevent multiple, but harmless warnings the code causes. Use of tidyverse would help clean this code up.
The p-values from all ten tests are sorted by strain in Table 2.
Table 2. Results of one sample t-tests inbred strains (Table 1) compared to average lifespan of the J:DO diversity outbred mouse strain from Jackson labs
| Strains | p-value | Strains | p-value | |
| 129S1/SvImJ | 0.8204 | C57BL/6J | 0.08623 | |
| A/J | 5.367e-08 | CBA/J | 1.601e-05 | |
| BALB/cByJ | 0.03527 | FVB/NJ | 0.3368 | |
| BUB/BnJ | 0.0004274 | P/J | 0.0007307 | |
| C3H/HeJ | 0.007479 | SWR/J | 0.2598 |
R-code for Fisher’s method
my_pValues <- c(0.08623, 1.601e-05, 0.3368, 0.0007307, 0.2598)
X_squared <- -2 * sum(log(my_pValues),base = exp(1))
myDf <- 2 * length(my_pValues)
combinedP <- pchisq(X_squared, myDf, lower.tail = FALSE)
result_table <- as.data.frame(rbind(X_squared, myDf, combinedP))
rownames(result_table) <- c("chi_sqr", "df", "P-value")
colnames(result_table) <- c("Value")
print(result_table)
and the R output
Value
chi_sqr 44.7017431931
df 16.0000000000
P-value 0.0001541831
With the small p-value, we would reject the null hypothesis of no difference in lifespan among the inbred strains compared to the genetically diverse DO strain of mice.
Limitations of ANOVA from sufficient statistics.
This was pretty easy, so it is worth asking — Why go through the bother of analyzing the raw data, why not just go to the summary statistics and run the calculator formula? First, the chief reason against the calculator formula and use of only sufficient statistics loses information about the individual values and therefore you have no access to the residuals. The residual of an observation is the difference between the original observation and the model prediction. The residuals are important for determining whether the model fits the data well and are, therefore, part of the toolkit that statisticians need to do proper data analysis. We will spend considerable time looking at residual patterns and it is an important aspect of doing statistics correctly.
Secondly, while it is possible to extend this approach to more complicated ANOVA problems like the two-way ANOVA (Cohen 2002), the statistical significance of the interaction term(s) calculated in this way are only approximate (the main effects are OK to interpret). Thus, ANOVA from sufficient statistics has its place when all you have is access to descriptive statistics, but its use is limited and not at all the preferred option for data analysis when the original, raw observations are in hand.
Questions
1. Calculate the one-way ANOVA for body weight of 47 female (F) and 97 male (M) cats (kilograms, dataset cats in MASS R package) from the following summary statistics.
| n | Mean | sd | |
| F | 47 | 2.36 | 0.274 |
| M | 97 | 2.9 | 0.468 |
2. Bonus: Load the cats data set (package MASS, loaded with Rcmdr) and run a one-way ANOVA using the aov() function via Rcmdr. Are the ANOVA from sufficient statistics the same as results from the direct ANOVA calculation? If not, why not.
Quiz Chapter 12.4
ANOVA from “sufficient statistics”