16.2 – Causation and Partial correlation
Introduction
Science driven by statistical inference and model building is largely motivated by the the drive to identify pathways of cause and effect linking events and phenomena observed all around us. (We first defined cause and effect in Chapter 2.4) The history of philosophy, from the works of Ancient Greece, China, Middle East and so on is rich in the language of cause and effect. From these traditions we have a number of ways to think of cause and effect, but for us it will be enough to review the logical distinction among three kinds of cause-effect associations:
- Necessary cause
- Sufficient cause
- Contributory cause
Here’s how the logic works. If A is a necessary cause of B, then the mere fact that B is present implies that A must also be present. However, that the presence of A does not imply that B will occur. If A is a sufficient cause of B, then the presence of A necessarily implies the presence of B. However, another cause C may alternatively cause B. Enter the contributory or related cause: A cause may be contributory if the presumed cause A (1) occurs before the effect B, and (2) changing A also changes B. A contributory cause does not need to be necessary nor must it be sufficient; contributory causes play a role in cause and effect.
Thus, following this long tradition of thinking about causality we have the mantra, “Correlation does not imply causation.” The exact phrase was written as early as emphasized by Karl Pearson who invented the statistic back in the late 1800s. This well-worn slogan deserves to be on T-shirts and bumper stickers*, and perhaps to be viewed as the single most important concept you can take from a course in philosophy/statistics. But in practice, we will always be tempted to stray from this guidance. The developments in genome-wide-association studies, or GWAS, are designed to look for correlations, as evidenced by statistical linkage analysis, between variation at one DNA base pair and presence/absence of disease or condition in humans and animal models. These are costly studies to do and in the end, the results are just that, evidence of associations (correlations), not proof of genetic cause and effect. We are less likely to be swayed by a correlation that is weak, but what about correlations that are large, even close to one? Is not the implication of high, statistically significant correlation evidence of causation? No, necessary, but not sufficient.
Note 1. A helpful review on causation in epidemiology is available from Parascandola and Weed (2001); see also Kleinberg and Hripcsak (2011). For more on “correlation does not imply causation”, try the Wikipedia entry. Obviously, researchers who engage in genome wide association studies are aware of these issues: see for example discussion by Hu et al (2018) on causal inference and GWAS.
Causal inference (Pearl 2009; Pearl and Mackenzie 2018), in brief, employs a model to explain the association between dependent and multiple, likely interrelated candidate causal variable, which is then subject to testing — is the model stable when the predictor variables are manipulated, when additional connections are considered (eg, predictor variable 1 covaries with one or more other predictor variables in the model). Wright’s path analysis, now included as one approach to Structural Equation Modeling, is used to relate equations (models) of variation in observed variables attributed to direct and indirect effects from predictor variables.
* And yes, a quick Google search reveals lots of bumper stickers and T-shirts available with the causation ![]() sentiment.
sentiment.
Spurious correlations
Correlation estimates should be viewed as hypotheses in the scientific sense of the meaning of hypotheses for putative cause-effect pairings. To drive the point home, explore the web site “Spurious Correlations” at https://www.tylervigen.com/spurious-correlations , which allows you to generate X-Y plots and estimate correlations among many different variables. Some of my favorite correlations from “Spurious Correlations” include (Table 1):
Table 1. Spurious correlations, https://www.tylervigen.com/spurious-correlations
| First variable | Second variable | Correlation |
| Divorce rate in Maine, USA | Per capita USA consumption of margarine | +0.993 |
| Honey producing bee colonies USA | Juvenile arrests for marijuana possession | -0.933 |
| Per capita USA consumption of mozzarella cheese | Civil engineering PhD awarded USA | +0.959 |
| Total number of ABA lawyers USA | Cost of red delicious apples | +0.879 |
These are some pretty strong correlations (cf. effect size discussion, Ch11.4 and 16.1), about as close to +1 as you can get. But really, do you think the amount of cheese that is consumed in the USA has anything to do with the number of PhD degrees awarded in engineering or that apple prices are largely set by the number of lawyers in the USA? Cause and effect implies there must also be some plausible mechanism, not just a strong correlation.
But that does NOT ALSO mean that a high correlation is meaningless. The primary reason a correlation cannot tell about causation is because of the problem (potentially) of an UNMEASURED variable (a confounding variable) being the real driving force (Fig 1).
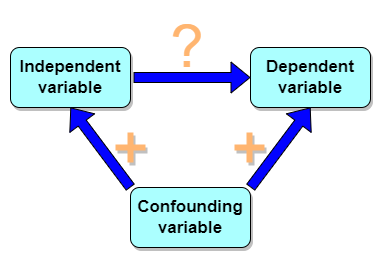
Figure 1. Unmeasured confounding variables influence association between independent and dependent variables, the characters or traits we are interested in.
Here’s a plot of running times for the best, fastest men and women runners for the 100-meter sprint. Running times over 100 meters of top athletes since the 1920s. The data are collated for you and presented at end of this page (scroll or click here).
Here’s a scatterplot (Fig 2).
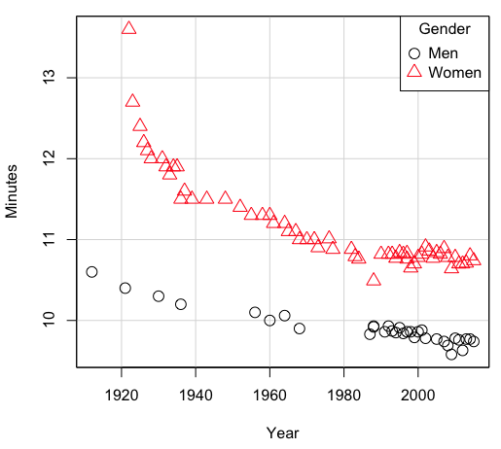
Figure 2. Running times over 100 meters of top athletes since the 1920s.
There’s clearly a negative correlation between years and running times. Is the rate of improvement in running times the same for men and women? Is the improvement linear? What, if any, are the possible confounding variables? Height? weight? Biomechanical differences? Society? Training? Genetics? … Performance enhancing drugs…?
If we measure potential confounding factors, we may be able to determine the strength of correlation between two variables that share variation with a third variable.
The partial correlation
There are several ways to work this problem. The partial correlation is a useful way to handle this problem, i.e., where a measured third variable is positively correlated with the two variable you are interested in.
![]()
Without formal mathematical proof presented, r12.3 is the correlation between variables 1 and 2 INDEPENDENT of any covariation with variable 3.
For our running data set, we have the correlation between women’s time for 100m over 9 decades, (r13 = -0.876), between men’s time for 100m over 9 decades (r23 = -0.952), and finally, the correlation we’re interested in, whether men’s and women’s times are correlated (r12 = +0.71). When we use the partial correlation, however, I get r12.3 = -0.819… much less than 0 and significantly different from zero. In other words, men’s and women’s times are not positively correlated independent of the correlation both share with the passage of time (decades)! The interpretation is that men are getting faster at a rate faster than women.
In conclusion, keep your head about you when you are doing analyses. You may not have the skills or knowledge to handle some problems (partial correlation), but you can think simply — why are two variables correlated? One causes the other to increase (or decrease) OR the two are both correlated with another variable.
Testing the partial correlation
Like our simple correlation, the partial correlation may be tested by a t-test, although modified to account for the number of pairwise correlations (Wetzels and Wagenmakers 2012). The equation for the t test statistic is now
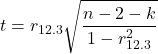
with k equal to the number of pairwise correlations and n – 2 – k degrees of freedom.
Examples
Lead exposure and low birth weigh. The data set is numbers of low birth weight births (< 2,500 g regardless of gestational age) and numbers of children with high levels of lead (10 or more micrograms of lead in a deciliter of blood) measured from their blood. Data used for 42 cities and towns of Rhode Island, United States of America (data at end of this page, scroll or click here to access the data).
A scatterplot of number of children with high lead is shown below (Fig 3).
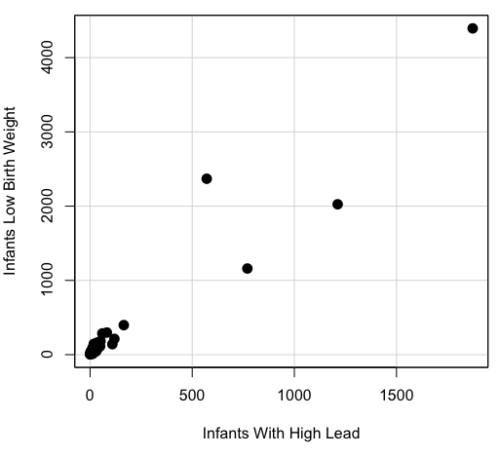
Figure 3. Scatterplot birth weight by lead exposure.
The product moment correlation was r = 0.961, t = 21.862, df = 40, p < 2.2e-16. So, at first blush looking at the scatterplot and the correlation coefficient, we conclude that there is a significant relationship between lead and low birth weight, right?
However, by the description of the data you should recall that counts were reported, not rates (eg, per 100,000 people). Clearly, population size varies among the cities and towns of Rhode Island. West Greenwich had 5085 people whereas Providence had 173,618. We should suspect that there is also a positive correlation between number of children born with low birth weight and numbers of children with high levels of lead. Indeed there are.
Correlation between Low Birth Weight and Population, r = 0.982
Correlation between High Lead levels and Population, r = 0.891
The question becomes, after removing the covariation with population size is there a linear association between high lead and low birth weight? One option is to calculate the partial correlation. To get partial correlations in Rcmdr, select
Statistics → Summaries → Correlation matrix
then select “partial” and select all three variables (Ctrl key) (Fig 4)
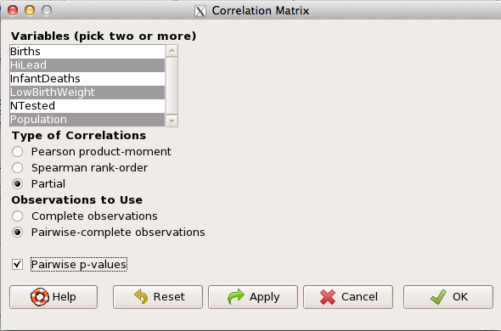
Figure 4. Screenshot Rcmdr partial correlation menu
Results are shown below.
partial.cor(leadBirthWeight[,c("HiLead","LowBirthWeight","Population")], tests=TRUE, use="complete")
Partial correlations:
HiLead LowBirthWeight Population
HiLead 0.00000 0.99181 -0.97804
LowBirthWeight 0.99181 0.00000 0.99616
Population -0.97804 0.99616 0.00000
Thus, after removing the covariation we conclude there is indeed a strong correlation between lead and low birth weights.
Note 2. Some verbiage about correlation tables (matrices). The matrix is symmetric and the information is repeated. I highlighted the diagonal in green. The upper triangle (red) is identical to the lower triangle (blue). When you publish such matrices, don’t publish both the upper and lower triangles; it’s also not necessary to publish the on-diagonal numbers, which are generally not of interest. Thus, the publishable matrix would be
Partial correlations:
LowBirthWeight Population
HiLead 0.99181 -0.97804
LowBirthWeight 0.99616
Another example
Do Democrats prefer cats? The question I was interested in, Do liberals really prefer cats, was inspired by a Time magazine 18 February 2014 article. I collated data on a separate but related question: Do states with more registered Democrats have more cat owners? The data set was compiled from three sources: 2010 USA Census, a 2011 Gallup poll about religious preferences, and from a data book on survey results of USA pet owners (data at end of this page, scroll or click here to access the data).
Note 3: This type of data set involves questions about groups, not individuals. We have access to aggregate statistics for groups (city, county, state, region), but not individuals. Thus, our conclusions are about groups and cannot be used to predict individual behavior, eg, knowing a person votes Green Party does not mean they necessarily share their home with a cat). See ecological fallacy.
This data set also demonstrates use of transformations of the data to improve fit of the data to statistical assumptions (normality, homoscedacity).
The variables, and their definitions, were:
ASDEMS = DEMOCRATS. Democrat advantage: the difference in registered Democrats compared to registered Republicans as a percentage; to improve the distribution qualities the arcsine transform was applied..
ASRELIG = RELIGION. Percent Religious from a Gallup poll who reported that Religion was “Very Important” to them. Also arcsine-transformed to improve normality and homoescedasticity (there you go, throwing $3 words around 🤠 ).
LGCAT = Number of pet cats, log10-transformed, estimated for USA states by survey, except Alaska and Hawaiʻi (not included in the survey by the American Veterinary Association).
LGDOG = Estimated number of pet dogs, log10-transformed for states, except Alaska and Hawaiʻi (not included in the survey by the American Veterinary Association).
LGIPC = Per capita income, log10-transformed.
LGPOP = Population size of each state, log10 transformed.
As always, begin with data exploration. All of the variables were right-skewed, so I applied data transformation functions as appropriate: log10 for the quantitative data and arc sine transform for the frequency variables. Because Democrat Advantage and Percent Religious variables were in percentages, the values were first divided by 100 to make frequencies, then the R function asin() was applied. All analyses were conducted on the transformed data, therefore conclusions apply to the transformed data.
Note 4: As we’ve discussed before, to relate the results to the original scales, back transformations would need to be run on any predictions. Back transformation for log10 would be power of ten; for the arcsine-transform the inverse of the arcsine would be used.
A scatter plot matrix (KMggplo2) plus histograms of the variables along the diagonals shows the results of the transforms and hints at the associations among the variables. A graphic like this one is called a trellis plot; a layout of smaller plots in a grid with the same (preferred) or at least similar axes. Trellis plots (Fig 5) are useful for finding the structure and patterns in complex data. Scanning across a row shows relationships between one variable with all of the others. For example, the first row Y-axis is for the ASDEMS variable; from left to right along the row we have, after the histogram, what look to be weak associations between ASDEMS and ASRELIG, LGCAT, LGDOG, and LGDOG.
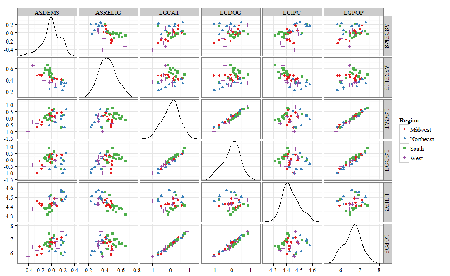
Figure 5. Trellis plot, correlations among variables.
A matrix of partial correlations was produced from the Rcmdr correlation call. Thus, to pick just one partial correlation, the association between DEMOCRATS and RELIGION (reported as “very important”) is negative (r = -0.45) and from the second matrix we retrieve the approximate P-value, unadjusted for the multiple comparisons problem, of p = 0.0024. We quickly move past this matrix to the adjusted P-values and confirm that this particular correlation is statistically significant even after correcting for multiple comparisons. Thus, there is a moderately strong negative correlation between those who reported that religion was very important to them and the difference between registered Democrats and Republicans in the 48 states. Because it is a partial correlation, we can conclude that this correlation is independent of all of the other included variables.
And what about our original question: Do Democrats prefer cats over dogs? The partial correlation after adjusting for all of the other correlated variables is small (r = 0.05) and not statistically different from zero (P-value greater than 5%).
Are there any interesting associations involving pet ownership in this data set? See if you can find it (hint: the correlation you are looking for is also in red).
Partial correlations:
ASDEMS ASRELIG LGCAT LGDOG LGIPC LGPOP
ASDEMS 0.0000 -0.4460 0.0487 0.0605 0.1231 -0.0044
ASRELIG -0.4460 0.0000 -0.2291 -0.0132 -0.4685 0.2659
LGCAT 0.0487 -0.2291 0.0000 0.2225 -0.1451 0.6348
LGDOG 0.0605 -0.0132 0.2225 0.0000 -0.6299 0.5953
LGIPC 0.1231 -0.4685 -0.1451 -0.6299 0.0000 0.6270
LGPOP -0.0044 0.2659 0.6348 0.5953 0.6270 0.0000
Raw P-values, Pairwise two-sided p-values:
ASDEMS ASRELIG LGCAT LGDOG LGIPC LGPOP ASDEMS 0.0024 0.7534 0.6965 0.4259 0.9772 ASRELIG 0.0024 0.1347 0.9325 0.0013 0.0810 LGCAT 0.7534 0.1347 0.1465 0.3473 <.0001 LGDOG 0.6965 0.9325 0.1465 <.0001 <.0001 LGIPC 0.4259 0.0013 0.3473 <.0001 <.0001 LGPOP 0.9772 0.0810 <.0001 <.0001 <.0001
Adjusted P-values, Holm’s method (Benjamini and Hochberg 1995)
ASDEMS ASRELIG LGCAT LGDOG LGIPC LGPOP ASDEMS 0.0241 1.0000 1.0000 1.0000 1.0000 ASRELIG 0.0241 1.0000 1.0000 0.0147 0.7293 LGCAT 1.0000 1.0000 1.0000 1.0000 <.0001 LGDOG 1.0000 1.0000 1.0000 <.0001 0.0002 LGIPC 1.0000 0.0147 1.0000 <.0001 <.0001 LGPOP 1.0000 0.7293 <.0001 0.0002 <.0001
A graph (Fig 6) to summarize the partial correlations: green lines indicate positive correlation, red lines show negative correlations; Strength of association indicated by the line thickness, with thicker lines corresponding to greater correlation.
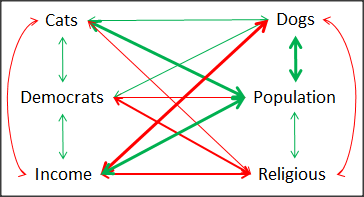
Figure 6. Causal paths among variables.
As you can see, partial correlation analysis is good for a few variables, but as the numbers increase it is difficult to make heads or tails out of the analysis. Better methods for working with these highly correlated data in what we call multivariate data analysis, for example Structural Equation Modeling or Path Analysis.
Questions
- Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
- Spurious correlations can be challenging to recognize, and, sometimes, they become part of a challenge to medicine to explain away. A classic spurious correlation is the correlation between rates of MMR vaccination and autism prevalence. Here’s a table of numbers for you
| Year | Herb Supplement Revenue, millions $ | Fertility rate per 1000 births, women age 35 and older+ | MMR per 100K children age 0 -5 | UFC revenue, millions $ | Autism Prevalence per 1000 |
|---|---|---|---|---|---|
| 2000 | 4225 | 47.7 | 179 | 6.7 | |
| 2001 | 4361 | 48.6 | 183 | 4.5 | |
| 2002 | 4275 | 49.9 | 190 | 8.7 | 6.6 |
| 2003 | 4146 | 52.6 | 196 | 7.5 | |
| 2004 | 4288 | 54.5 | 199 | 14.3 | 8 |
| 2005 | 4378 | 55.5 | 197 | 48.3 | |
| 2006 | 4558 | 56.9 | 198 | 180 | 9 |
| 2007 | 4756 | 57.6 | 204 | 226 | |
| 2008 | 4800 | 56.7 | 202 | 275 | 11.3 |
| 2009 | 5037 | 56.1 | 201 | 336 | |
| 2010 | 5049 | 56.1 | 209 | 441 | 14.4 |
| 2011 | 5302 | 57.5 | 212 | 437 | |
| 2012 | 5593 | 58.7 | 216 | 446 | 14.5 |
| 2013 | 6033 | 59.7 | 220 | 516 | |
| 2014 | 6441 | 61.6 | 224 | 450 | 16.8 |
| 2015 | 6922 | 62.8 | 222 | 609 | |
| 2016 | 7452 | 64.1 | 219 | 666 | 18.5 |
| 2017 | 8085 | 63.9 | 213 | 735 | |
| 2018 | 65.3 | 220 | 800 | 25 |
- Make scatterplots of autism prevalence vs
- Herb supplement revenue
- Fertility rate
- MMR vaccination
- UFC revenue
- Calculate and test correlations between autism prevalence vs
- Herb supplement revenue
- Fertility rate
- MMR vaccination
- UFC revenue
- Interpret the correlations — is there any clear case for autism vs MMR?
- What additional information is missing from Table 2? Add that missing variable and calculate partial correlations for autism prevalence vs
- Herb supplement revenue
- Fertility rate
- MMR vaccination
- UFC revenue
- Do a little research: What are some reasons for increase in autism prevalence? What is the consensus view about MMR vaccine and risk of autism?
Quiz Chapter 16.2
Causation and Partial correlation
Data sets
Data used in this page, 100 meter running times since 1900.
| Year | Men | Women |
|---|---|---|
| 1912 | 10.6 | |
| 1913 | ||
| 1914 | ||
| 1915 | ||
| 1916 | ||
| 1917 | ||
| 1918 | ||
| 1919 | ||
| 1920 | ||
| 1921 | 10.4 | |
| 1922 | 13.6 | |
| 1923 | 12.7 | |
| 1924 | ||
| 1925 | 12.4 | |
| 1926 | 12.2 | |
| 1927 | 12.1 | |
| 1928 | 12 | |
| 1929 | ||
| 1930 | 10.3 | |
| 1931 | 12 | |
| 1932 | 11.9 | |
| 1933 | 11.8 | |
| 1934 | 11.9 | |
| 1935 | 11.9 | |
| 1936 | 10.2 | 11.5 |
| 1937 | 11.6 | |
| 1938 | ||
| 1939 | 11.5 | |
| 1940 | ||
| 1941 | ||
| 1942 | ||
| 1943 | 11.5 | |
| 1944 | ||
| 1945 | ||
| 1946 | ||
| 1947 | ||
| 1948 | 11.5 | |
| 1949 | ||
| 1950 | ||
| 1951 | ||
| 1952 | 11.4 | |
| 1953 | ||
| 1954 | ||
| 1955 | 11.3 | |
| 1956 | 10.1 | |
| 1957 | ||
| 1958 | 11.3 | |
| 1959 | ||
| 1960 | 10 | 11.3 |
| 1961 | 11.2 | |
| 1962 | ||
| 1963 | ||
| 1964 | 10.06 | 11.2 |
| 1965 | 11.1 | |
| 1966 | ||
| 1967 | 11.1 | |
| 1968 | 9.9 | 11 |
| 1969 | ||
| 1970 | 11 | |
| 1972 | 10.07 | 11 |
| 1973 | 10.15 | 10.9 |
| 1976 | 10.06 | 11.01 |
| 1977 | 9.98 | 10.88 |
| 1978 | 10.07 | 10.94 |
| 1979 | 10.01 | 10.97 |
| 1980 | 10.02 | 10.93 |
| 1981 | 10 | 10.9 |
| 1982 | 10 | 10.88 |
| 1983 | 9.93 | 10.79 |
| 1984 | 9.96 | 10.76 |
| 1987 | 9.83 | 10.86 |
| 1988 | 9.92 | 10.49 |
| 1989 | 9.94 | 10.78 |
| 1990 | 9.96 | 10.82 |
| 1991 | 9.86 | 10.79 |
| 1992 | 9.93 | 10.82 |
| 1993 | 9.87 | 10.82 |
| 1994 | 9.85 | 10.77 |
| 1995 | 9.91 | 10.84 |
| 1996 | 9.84 | 10.82 |
| 1997 | 9.86 | 10.76 |
| 1998 | 9.86 | 10.65 |
| 1999 | 9.79 | 10.7 |
| 2000 | 9.86 | 10.78 |
| 2001 | 9.88 | 10.82 |
| 2002 | 9.78 | 10.91 |
| 2003 | 9.93 | 10.86 |
| 2004 | 9.85 | 10.77 |
| 2005 | 9.77 | 10.84 |
| 2006 | 9.77 | 10.82 |
| 2007 | 9.74 | 10.89 |
| 2008 | 9.69 | 10.78 |
| 2009 | 9.58 | 10.64 |
| 2010 | 9.78 | 10.78 |
| 2011 | 9.76 | 10.7 |
| 2012 | 9.63 | 10.7 |
| 2013 | 9.77 | 10.71 |
| 2014 | 9.77 | 10.8 |
| 2015 | 9.74 | 10.74 |
| 2016 | 9.8 | 10.7 |
| 2017 | 9.82 | 10.71 |
| 2018 | 9.79 | 10.85 |
| 2019 | 9.76 | 10.71 |
| 2020 | 9.86 | 10.85 |
| 2021 | 9.77 | 10.54 |
| 2022 | 9.76 | 10.62 |
| 2023 | 9.83 | 10.65 |
| 2024 | 9.77 | 10.72 |
Data used in this page, birth weight by lead exposure
| CityTown | Core | Population | NTested | HiLead | Births | LowBirthWeight | InfantDeaths |
|---|---|---|---|---|---|---|---|
| Barrington | n | 16819 | 237 | 13 | 785 | 54 | 1 |
| Bristol | n | 22649 | 308 | 24 | 1180 | 77 | 5 |
| Burrillville | n | 15796 | 177 | 29 | 824 | 44 | 8 |
| Central Falls | y | 18928 | 416 | 109 | 1641 | 141 | 11 |
| Charlestown | n | 7859 | 93 | 7 | 408 | 22 | 1 |
| Coventry | n | 33668 | 387 | 20 | 1946 | 111 | 7 |
| Cranston | n | 79269 | 891 | 82 | 4203 | 298 | 20 |
| Cumberland | n | 31840 | 381 | 16 | 1669 | 98 | 8 |
| East Greenwich | n | 12948 | 158 | 3 | 598 | 41 | 3 |
| East Providence | n | 48688 | 583 | 51 | 2688 | 183 | 11 |
| Exeter | n | 6045 | 73 | 2 | 362 | 6 | 1 |
| Foster | n | 4274 | 55 | 1 | 208 | 9 | 0 |
| Glocester | n | 9948 | 80 | 3 | 508 | 32 | 5 |
| Hopkintown | n | 7836 | 82 | 5 | 484 | 34 | 3 |
| Jamestown | n | 5622 | 51 | 14 | 215 | 13 | 0 |
| Johnston | n | 28195 | 333 | 15 | 1582 | 102 | 6 |
| Lincoln | n | 20898 | 238 | 20 | 962 | 52 | 4 |
| Little Compton | n | 3593 | 48 | 3 | 134 | 7 | 0 |
| Middletown | n | 17334 | 204 | 12 | 1147 | 52 | 7 |
| Narragansett | n | 16361 | 173 | 10 | 728 | 42 | 3 |
| Newport | y | 26475 | 356 | 49 | 1713 | 113 | 7 |
| New Shoreham | n | 1010 | 11 | 0 | 69 | 4 | 1 |
| North Kingstown | n | 26326 | 378 | 20 | 1486 | 76 | 7 |
| North Providence | n | 32411 | 311 | 18 | 1679 | 145 | 13 |
| North Smithfield | n | 10618 | 106 | 5 | 472 | 37 | 3 |
| Pawtucket | y | 72958 | 1125 | 165 | 5086 | 398 | 36 |
| Portsmouth | n | 17149 | 206 | 9 | 940 | 41 | 6 |
| Providence | y | 173618 | 3082 | 770 | 13439 | 1160 | 128 |
| Richmond | n | 7222 | 102 | 6 | 480 | 19 | 2 |
| Scituate | n | 10324 | 133 | 6 | 508 | 39 | 2 |
| Smithfield | n | 20613 | 211 | 5 | 865 | 40 | 4 |
| South Kingstown | n | 27921 | 379 | 35 | 1330 | 72 | 10 |
| Tiverton | n | 15260 | 174 | 14 | 516 | 29 | 3 |
| Warren | n | 11360 | 134 | 17 | 604 | 42 | 1 |
| Warwick | n | 85808 | 973 | 60 | 4671 | 286 | 26 |
| Westerly | n | 22966 | 140 | 11 | 1431 | 85 | 7 |
| West Greenwich | n | 5085 | 68 | 1 | 316 | 15 | 0 |
| West Warwick | n | 29581 | 426 | 34 | 2058 | 162 | 17 |
| Woonsoket | y | 43224 | 794 | 119 | 2872 | 213 | 22 |
Data in this page, Do Democrats prefer cats?
Chapter 16 contents
- Introduction
- Product moment correlation
- Causation and Partial correlation
- Data aggregation and correlation
- Spearman and other correlations
- Instrument reliability and validity
- Similarity and Distance
- References and suggested readings
15.3 – Wilcoxon signed rank test
Introduction
When experimental units are repeated or paired, they lack independence and evaluating any difference between paired groups without accounting for the association between the repeat measures or between the pairs of related subjects would lead to incorrect inferences. A familiar pairing of experimental units occurs in clinical observational research in which control subjects and treatment subjects are matched by many characteristics.
In such cases, the parametric paired t-test would be used to evaluate inferences about the differences between repeat measures or between treatment and matched control subjects for some measured outcome. A nonparametric alternative to the paired t-test is the Wilcoxon signed rank test, also called paired Wilcoxon test.
Another common example of paired sampling units would be that individuals are measured more than once for the same character or feature. For example, Chapter 10.3 we presented results of running pace in minutes to complete the race of 15 women for repeated trials (different years) at a 5K race held annually in Honolulu (Table 1).
Table 1. 5K repeat measures running data.
| ID | First race | Second race |
|---|---|---|
| 1 | 15.28 | 15.61 |
| 2 | 11.22 | 11.19 |
| 3 | 8.80 | 9.14 |
| 4 | 8.88 | 5.46 |
| 5 | 9.81 | 10.50 |
| 6 | 6.12 | 5.69 |
| 7 | 8.31 | 8.71 |
| 8 | 6.26 | 7.42 |
| 9 | 17.16 | 16.41 |
| 10 | 16.23 | 15.82 |
| 11 | 5.90 | 7.12 |
| 12 | 8.31 | 10.48 |
| 13 | 5.93 | 8.64 |
| 14 | 10.54 | 5.99 |
| 15 | 9.53 | 8.69 |
Data set introduced in Chapter 10.3
To get the paired Wilcoxon test in R Commander, select Rcmdr: Statistics → Nonparametric tests → Paired Wilcoxon Test
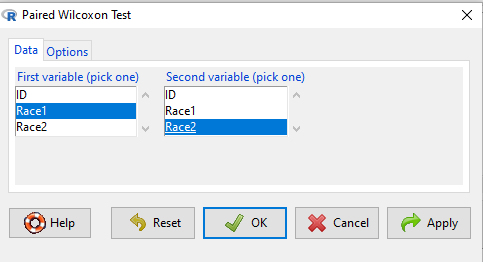
Figure 1. R Commander paired Wilcoxon test menu (aka Wilcoxon signed rank sum test). Rcmdr version 2.7.
Select first variable (e.g., Race1), second variable (e.g., Race2). Next, select Options tab and set null hypothesis. Accept defaults (Fig. 2).
Figure 2. R Commander Options, select null hypothesis.
For the nonparametric paired Wilcoxon test we choose among the options to set our conditions; from the context menu a two-tailed test and we elect to use the defaults for the tests and calculations of P-values.
The results, copied from the Output window, are shown below. The calculation of the Wilcoxon test statistic (V) is straightforward, involving summing the ranks. Obtaining the P-value of the test of the null is a bit more involved as it depends on permutations of all possible combinations of differences. For us, R will do nicely with the details, and we just need to check the P-value.
with(repeat15_banana5K, wilcox.test(Race.1, Race.2, alternative='two.sided', paired=TRUE)) # median difference [1] -0.3313126 Wilcoxon signed rank exact test data: Race.1 and Race.2 V = 58, p-value = 0.9341 alternative hypothesis: true location shift is not equal to 0
End R output
Here, we see that the median difference is small (-0.33), and the associated P-value is 0.93. Thus, we failed to reject the null hypothesis and should conclude that there was no difference in median running pace during the first and second trials.
Note that this is the same general conclusion we got when we ran a paired t-test on this data set: no difference between day one and day two.
R code
example.ch10.3 <- read.table(header=TRUE, text = " ID Race1 Race2 1 15.28 15.61 2 11.22 11.19 3 8.80 9.14 4 8.88 5.46 5 9.81 10.50 6 6.12 5.69 7 8.31 8.71 8 6.26 7.42 9 17.16 16.41 10 16.23 15.82 11 5.90 7.12 12 8.31 10.48 13 5.93 8.64 14 10.54 5.99 15 9.53 8.69 ")
Questions
- This question lists all fourteen statistical tests we have been introduced to so far
a. Mark yes or no as to whether or not the test is a parametric test
b. Identify the nonparametric test(s) with their equivalent parametric test(s). If there are no equivalency, simply write “none.”
|
|
Parametric test? |
If nonparametric, write the number(s) of the tests that the nonparametric test serves as an alternate for |
|
1. ANOVA by ranks |
|
|
|
2. Bartlett Test |
|
|
|
3. Chi-squared contingency table |
|
|
|
4. Chi-squared goodness of fit |
|
|
|
5. Fisher Exact test |
|
|
|
6. Independent sample T-test |
|
|
|
7. Kruskal-Walis test |
|
|
|
8. Levene Test |
|
|
|
9. One-sample T-test |
|
|
|
10. One-way ANOVA |
|
|
|
11. Paired T-test |
|
|
|
12. Shapiro-Wilks test |
|
|
|
13. Tukey posthoc comparisons |
|
|
|
14. Welch’s test |
|
|
Quiz Chapter 15.3
Wilcoxon Signed Rank Test
Chapter 15 contents
- Introduction
- Kruskal-Wallis and ANOVA by ranks
- Wilcoxon Rank Sum Test
- Wilcoxon signed rank test
- References and suggested readings
15.1 – Kruskal-Wallis and ANOVA by ranks
Introduction
When the data are NOT normally distributed OR When the variances in the different samples are unequal, one option is to opt for a nonparametric alternative use the Kruskal-Wallis test. The test is the same as an ANOVA on ranks. The tests are used to test whether or not groups can be viewed as having come from the same sample distribution. This is a terse page with emphasis on examples.
It is known that the an ANOVA on ranks (in blue) of the original data (in black) will yield the same results.
Kruskal-Wallis
Rcmdr: Statistics → Nonparametric test → Kruskal-Wallis test…
Rcmdr Output of Kruskal-Wallis test
tapply(Pop_data$Stuff, Pop_data$Pop, median, na.rm=TRUE) Pop1 Pop2 Pop3 Pop4 146 90 122 347 kruskal.test(Stuff ~ Pop, data=Pop_data) Kruskal-Wallis rank sum test data: Stuff by Pop Kruskal-Wallis chi-squared = 25.6048, df = 3, p-value = 1.154e-05
End of R output
So, we reject — “provisionally, of course! — the null hypothesis, right?
Compare parametric test and alternative non-parametric test
Let’s compare the nonparametric test results to those from an analysis of the ranks (ANOVA of ranks).
To get the ranks in R Commander (example.15.1 data set is available at bottom of this page; scroll down or click here).
Rcmdr: Data → Manage variables in active data set → Compute new variable …
The command for ranks is…. wait for it …. rank(). In the popup menu box, name the new variable (Ranks) and in the Expression to compute box enter rank(Values).
Figure 1. Screenshot Rcmdr menu create new variable.
Note 1: Not a good idea to name an object, Ranks, that’s similar to a function name in R, rank.
And the R code is simply
example.15.1$Ranks <- with(example.15.1, rank(Values))
Note 2: The object example.15.1$Ranks adds our new variable to our data frame.
That’s one option, rank across the entire data set. Another option would be to rank within groups.
R code
example.15.1$xRanks <- ave(Values, Population, FUN=rank)
Note 3: The ave() function averages within subsets of the data and applies whatever summary function (FUN) you choose. In this case we used rank. Alternative approaches could use split or lapply or variations of dplyr. ave() is in the base package and at least in this case is simply to use to solve our rank within groups problem. xRanks would then be added to the existing data frame.
Here are the results of ranking within groups (Table 1).
Table 1. Original and ranked data set.
| Population 1 | Rank1 | Population 2 | Rank2 | Population 3 | Rank3 | Population 4 | Rank4 |
| 105 | 11.5 | 100 | 9 | 130 | 17.5 | 310 | 33 |
| 132 | 19 | 65 | 4 | 95 | 7 | 302 | 32 |
| 156 | 22 | 60 | 2.5 | 100 | 9 | 406 | 38 |
| 198 | 29 | 125 | 16 | 124 | 15 | 325 | 34 |
| 120 | 13.5 | 80 | 5.5 | 120 | 13.5 | 298 | 31 |
| 196 | 28 | 140 | 21 | 180 | 26 | 412 | 39.5 |
| 175 | 24 | 50 | 1 | 80 | 5.5 | 385 | 39.5 |
| 180 | 26 | 180 | 26 | 210 | 30 | 329 | 35 |
| 136 | 20 | 60 | 2.5 | 100 | 9 | 375 | 37 |
| 105 | 115 | 130 | 17.5 | 170 | 23 | 365 | 36 |
Question 1. Which do you choose, rank across groups (Ranks) or rank within groups (xRanks)? Recall that this example began with a nonparametric alternative to the one-way ANOVA, and we were testing the null hypothesis that the group means were the same
![]()
Answer 1. Rank the entire data set, ignoring the groups. The null hypothesis here is that there is no difference in median ranks among the groups. Ranking within groups simply shuffles observations within the group. This is basically the same thing as running Kruskal-Wallis test.
Run the one-way ANOVA, now on the Ranked variable. The ANOVA table is summarized Table 2.
Table 2. ANOVA table on ranked variable.
| Source | DF | SS | MS | F | P† |
| Population | 3 | 3495.1 | 1165.0 | 22.94 | < 0.001 |
| Error | 36 | 1828.0 | 50.8 | ||
| Total | 39 | 5323.0 |
Note 4: † The exact p-value returned by R was 0.0000000178. This level of precision is a bit suspect given that calculations of p-values are subject to bias too, like any estimate. Thus, some advocate to report p-values to three significant figures and if less than 0.001, report as shown in this table. Occasionally, you may see P = 0.000 written in a journal article. This is a definite no-no; p-values are estimates of the probability of getting results more extreme then our results and the null hypothesis holds. It’s an estimate, not certainty; p-values cannot equal zero.
So, how do you choose between the parametric ANOVA and the non-parametric Kruskal-Wallis (ANOVA by Ranks) test? Think like a statistician — It is all about the type I error rate and potential bias of a statistical test. The purpose of statistics is to help us separate real effects from random chance differences. If we are employing the NHST approach, then we must consider our chance that we are committing either a Type I error or a Type II error, and conservative tests, eg, tests based on comparing medians and not means, implies an increased chance of committing Type II errors.
Questions
- Saying that nonparametric tests make fewer assumptions about the data should not be interpreted that they make no assumptions about the data. Thinking about our discussions about experimental design and our discussion about test assumptions, what assumptions must hold regardless of the statistical test used?
- Go ahead and carry out the one-way ANOVA on the within group ranks (
xRanks). What’s the p-value from the ANOVA? - One could take the position that only nonparametric alternative tests should be employed in place of parametric tests, in part because they make fewer assumptions about the data. Why is this position unwarranted?
Quiz Chapter 15.1
Kruskal-Wallis and ANOVA by ranks
Data used in this page
Dataset for Kruskal-Wallis test
| Population | Values |
|---|---|
| Pop1 | 105 |
| Pop1 | 132 |
| Pop1 | 156 |
| Pop1 | 198 |
| Pop1 | 120 |
| Pop1 | 196 |
| Pop1 | 175 |
| Pop1 | 180 |
| Pop1 | 136 |
| Pop1 | 105 |
| Pop2 | 100 |
| Pop2 | 65 |
| Pop2 | 60 |
| Pop2 | 125 |
| Pop2 | 80 |
| Pop2 | 140 |
| Pop2 | 50 |
| Pop2 | 180 |
| Pop2 | 60 |
| Pop2 | 130 |
| Pop3 | 130 |
| Pop3 | 95 |
| Pop3 | 100 |
| Pop3 | 124 |
| Pop3 | 120 |
| Pop3 | 180 |
| Pop3 | 80 |
| Pop3 | 210 |
| Pop3 | 100 |
| Pop3 | 170 |
| Pop4 | 310 |
| Pop4 | 302 |
| Pop4 | 406 |
| Pop4 | 325 |
| Pop4 | 298 |
| Pop4 | 412 |
| Pop4 | 385 |
| Pop4 | 329 |
| Pop4 | 375 |
| Pop4 | 365 |
Chapter 15 contents
- Introduction
- Kruskal-Wallis and ANOVA by ranks
- Wilcoxon Rank Sum Test
- Wilcoxon signed rank test
- References and suggested readings
20.10 – Growth equations and dose response calculations
Introduction
In biology, growth may refer to increase in cell number, change in size of an individual across development, or increase of number of individuals in a population over time. Nonlinear, time-series, several models proposed to fit growth data, including the Gompertz, logistic, and the von Bertalanffy. These models fit many S-shaped growth curves. These models are special cases of generalized linear models, also called Richard curves.
Growth example
This page describes how to use R to analyze growth curve data sets.
| Hours | Abs |
|---|---|
| 0.000 | 0.002207 |
| 0.274 | 0.010443 |
| 0.384 | 0.033688 |
| 0.658 | 0.063257 |
| 0.986 | 0.111848 |
| 1.260 | 0.249240 |
| 1.479 | 0.416236 |
| 1.699 | 0.515578 |
| 1.973 | 0.572632 |
| 2.137 | 0.589528 |
| 2.466 | 0.619091 |
| 2.795 | 0.608486 |
| 3.123 | 0.621136 |
| 3.671 | 0.616850 |
| 4.110 | 0.614689 |
| 4.548 | 0.614643 |
| 5.151 | 0.612465 |
| 5.534 | 0.606082 |
| 5.863 | 0.603933 |
| 6.521 | 0.595407 |
| 7.068 | 0.589006 |
| 7.671 | 0.578372 |
| 8.164 | 0.567749 |
| 8.877 | 0.559217 |
| 9.644 | 0.546451 |
| 10.466 | 0.537907 |
| 11.233 | 0.537826 |
| 11.890 | 0.529300 |
| 12.493 | 0.516551 |
| 13.205 | 0.505905 |
| 14.082 | 0.491013 |
Key to parameter estimates: y0 is the lag, mumax is the growth rate, and K is the asymptotic stationary growth phase. The spline function does not return an estimate for K.
R code
require(growthrates) #Enter the data. Replace these example values with your own #time variable (Hours) Hours <- c(0.000, 0.274, 0.384, 0.658, 0.986, 1.260, 1.479, 1.699, 1.973, 2.137, 2.466, 2.795, 3.123, 3.671, 4.110, 4.548, 5.151, 5.534, 5.863, 6.521, 7.068, 7.671, 8.164, 8.877, 9.644, 10.466, 11.233, 11.890, 12.493, 13.205, 14.082) #absorbance or concentration variable (Abs) Abs <- c(0.002207, 0.010443, 0.033688, 0.063257, 0.111848, 0.249240, 0.416236, 0.515578, 0.572632, 0.589528, 0.619091, 0.608486, 0.621136, 0.616850, 0.614689, 0.614643, 0.612465, 0.606082, 0.603933, 0.595407, 0.589006, 0.578372, 0.567749, 0.559217, 0.546451, 0.537907, 0.537826, 0.529300, 0.516551, 0.505905, 0.491013)
#Make a dataframe and check the data; If error, then check that variables have equal numbers of entries Yeast <- data.frame(Hours,Abs); Yeast
#Obtain growth parameters from fit of a parametric growth model
#First, try some reasonable starting values
p <- c(y0 = 0.001, mumax = 0.5, K = 0.6)
model.par <- fit_growthmodel(FUN = grow_logistic, p = p, Hours, Abs, method=c("L-BFGS-B"))
summary(model.par)
coef(model.par)
#Obtain growth parameters from fit of a nonparametric smoothing spline model.npar <- fit_spline(Hours,Abs) summary(model.npar) coef(spline.md)
#Make plots par(mfrow = c(2, 1)) plot(Yeast, ylim=c(0,1), cex=1.5,pch=16, main="Parametric Nonlinear Growth Model", xlab="Hours", ylab="Abs") lines(model.par, col="blue", lwd=2) plot(model.npar, ylim=c(0,1), lwd=2, main="Nonparametric Spline Fit", xlab="Hours", ylab="Abs")
Results from example code
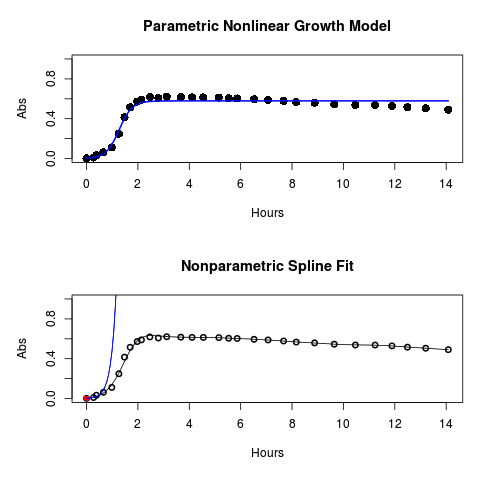
Figure 1. Top: Parametric Nonlinear Growth Model; Bottom: Nonparametric Spline Fit
LD50
In toxicology, the dose of a pathogen, radiation, or toxin required to kill half the members of a tested population of animals or cells is called the lethal dose, 50%, or LD50. This measure is also known as the lethal concentration, LC50, or properly after a specified test duration, the LCt50 indicating the lethal concentration and time of exposure. LD50 figures are frequently used as a general indicator of a substance’s acute toxicity. A lower LD50 is indicative of increased toxicity.
The point at which 50% response of studied organisms to range of doses of a substance (e.g., agonist, antagonist, inhibitor, etc.) to any response, from change in behavior or life history characteristics up to and including death can be described by the methods described in this chapter. The procedures outlined below assume that there is but one inflection point, i.e., an “s-shaped” curve, either up or down; if there are more than one inflection points, then the logistic equations described will not fit the data well and other choices need to be made (see Di Veroli et al 2015). We will use the drc package (Ritz et al 2015).
Example
First we’ll work through use of R. We’ll follow up with how to use Solver in Microsoft Excel.
After starting R, load the drc library.
library(drc)
Consider some hypothetical 24-hour survival data for yeast exposed to salt solutions. Let resp equal the variable for frequency of survival (e.g., calculated from OD660 readings) and NaCl equal the millimolar (mm) salt concentrations or doses.
At the R prompt type
resp <- c(1,1,1,.9,.7,.3,.4,.2,0,0,0) NaCl=seq(0,1000,100) #Confirm sequence was correctly created; alternatively, enter the values. NaCl [1] 0 100 200 300 400 500 600 700 800 900 1000 #Make a plot plot(NaCl,resp,pch=19,cex=1.2,col="blue",xlab="NaCl [mm]",ylab="Survival frequency")
And here is the plot (Fig. 2).
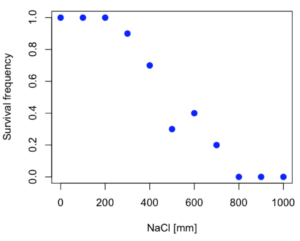
Figure 2. Hypothetical data set, survival of yeast in different salt concentrations.
Note the sigmoidal “S” shape — we’ll need an logistic equation to describe the relationship between survival of yeast and NaCl doses.
The equation for the four parameter logistic curve, also called the Hill-Slope model, is
![]()
where c is the parameter for the lower limit of the response, d is the parameter for the upper limit of the response, e is the relative EC50, the dose fitted half-way between the limits c and d, and b is the relative slope around the EC50. The slope, b, is also known as the Hill slope. Because this experiment included a dose of zero, a three parameter logistic curve would be appropriate. The equation simplifies to
![]()
EC50 from 4 parameter model
Let’s first make a data frame
dose <- data.frame(NaCl, resp)
Then call up a function, drm, from the drc library and specify the model as the four parameter logistic equation, specified as LL.4(). We follow with a call to the summary command to retrieve output from the drm function. Note that the four-parameter logistic equation
model.dose1 = drm(dose,fct=LL.4()) summary(model.dose1)
And here is the R output.
Model fitted: Log-logistic (ED50 as parameter) (4 parms) Parameter estimates: Estimate Std. Error t-value p-value b:(Intercept) 3.753415 1.074050 3.494636 0.0101 c:(Intercept) -0.084487 0.127962 -0.660251 0.5302 d:(Intercept) 1.017592 0.052460 19.397441 0.0000 e:(Intercept) 492.645128 47.679765 10.332373 0.0000 Residual standard error: 0.0845254 (7 degrees of freedom)
The EC50, technically the LD50 because the data were for survival, is the value of e: 492.65 mM NaCl.
You should always plot the predicted line from your model against the real data and inspect the fit.
At the R prompt type
plot(model.dose1, log="",pch=19,cex=1.2,col="blue",xlab="NaCl [mm]",ylab="Survival frequency")
As long as the plot you made in earlier steps is still available, R will add the line specified in the lines command. Here is the plot with the predicted logistic line displayed (Fig. 3).
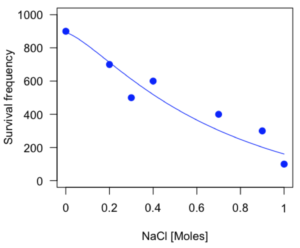
Figure 3. Logistic curve added to Figure 1 plot.
While there are additional steps we can take to decide is the fit of the logistic curve was good to the data, visual inspection suggests that indeed the curve fits the data reasonably well.
More work to do
Because the EC50 calculations are estimates, we should also obtain confidence intervals. The drc library provides a function called ED which will accomplish this. We can also ask what the survival was at 10% and 90% in addition to 50%, along with the confidence intervals for each.
At the R prompt type
ED(model.dose1,c(10,50,90), interval="delta")
And the output is shown below.
Estimated effective doses (Delta method-based confidence interval(s)) Estimate Std. Error Lower Upper 1:10 274.348 38.291 183.803 364.89 1:50 492.645 47.680 379.900 605.39 1:90 884.642 208.171 392.395 1376.89
Thus, the 95% confidence interval for the EC50 calculated from the four parameter logistic curve was between the lower limit of 379.9 and an upper limit of 605.39 mm NaCl.
EC50 from 3 parameter model
Looking at the summary output from the four parameter logistic function we see that the value for c was -0.085 and p-value was 0.53, which suggests that the lower limit was not statistically different from zero. We would expect this given that the experiment had included a control of zero mm added salt. Thus, we can explore by how much the EC50 estimate changes when the additional parameter c is no longer estimated by calculating a three parameter model with LL.3().
model.dose2 = drm(dose,fct=LL.3()) summary(model.dose2)
R output follows.
Model fitted: Log-logistic (ED50 as parameter) with lower limit at 0 (3 parms) Parameter estimates: Estimate Std. Error t-value p-value b:(Intercept) 4.46194 0.76880 5.80378 4e-04 d:(Intercept) 1.00982 0.04866 20.75272 0e+00 e:(Intercept) 467.87842 25.24633 18.53253 0e+00 Residual standard error: 0.08267671 (8 degrees of freedom)
The EC50 is the value of e: 467.88 mM NaCl.
How do the four and three parameter models compare? We can rephrase this as as statistical test of fit; which model fits the data better, a three parameter or a four parameter model?
At the R prompt type
anova(model.dose1, model.dose2)
The R output follows.
1st mode fct: LL.3() 2nd model fct: LL.4() ANOVA table ModelDf RSS Df F value p value 2nd model 8 0.054684 1st model 7 0.050012 1 0.6539 0.4453
Because the p-value is much greater than 5% we may conclude that the fit of the four parameter model was not significantly better than the fit of the three parameter model. Thus, based on our criteria we established in discussions of model fit in Chapter 16 – 18, we would conclude that the three parameter model is the preferred model.
The plot below now includes the fit of the four parameter model (red line) and the three parameter model (green line) to the data (Fig. 4).
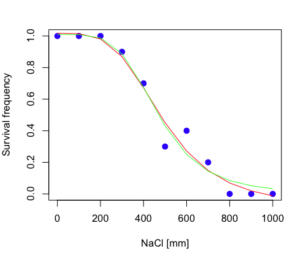
Figure 4. Four parameter (red) and three parameter (green) logistic models fitted to data.
The R command to make this addition to our active plot was
lines(dose,predict(model.dose2, data.frame(x=dose)),col="green")
We continue with our analysis of the three parameter model and produce the confidence intervals for the EC50 (modify the ED() statement above for model.dose2 in place of model.dose1).
Estimated effective doses (Delta method-based confidence interval(s)) Estimate Std. Error Lower Upper 1:10 285.937 33.154 209.483 362.39 1:50 467.878 25.246 409.660 526.10 1:90 765.589 63.026 620.251 910.93
Thus, the 95% confidence interval for the EC50 calculated from the three parameter logistic curve was between the lower limit of 409.7 and an upper limit of 526.1 mm NaCl. The difference between upper and lower limits was 116.4 mm NaCl, a smaller difference than the interval calculated for the 95% confidence intervals from the four parameter model (225.5 mm NaCl). This demonstrates the estimation trade-off: more parameters to estimate reduces the confidence in any one parameter estimate.
Additional notes of EC50 calculations
Care must be taken that the model fits the data well. What if we did not have observations throughout the range of the sigmoidal shape? We can explore this by taking a subset of the data.
dd = dose[1:6,]
Here, all values after dose 500 were dropped
dd resp dose 1 1.0 0 2 1.0 100 3 1.0 200 4 0.9 300 5 0.7 400 6 0.3 500
and the plot does not show an obvious sigmoidal shape (Fig. 5).
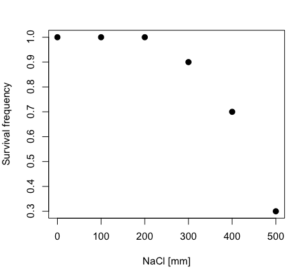
Figure 5. Plot of reduced data set.
We run the three parameter model again, this time on the subset of the data.
model.dosedd = drm(dd,fct=LL.3()) summary(model.dosedd)
Output from the results are
Model fitted: Log-logistic (ED50 as parameter) with lower limit at 0 (3 parms) Parameter estimates: Estimate Std. Error t-value p-value b:(Intercept) 6.989842 0.760112 9.195801 0.0027 d:(Intercept) 0.993391 0.014793 67.153883 0.0000 e:(Intercept) 446.882542 5.905728 75.669344 0.0000 Residual standard error: 0.02574154 (3 degrees of freedom)
Conclusion? The estimate is different, but only just so, 447 vs. 468 mm NaCl.
Thus, within reason, the drc function performs well for the calculation of EC50. Not all tools available to the student will do as well.
NLopt and nloptr
draft
Free open source library for nonlinear optimization.
Steven G. Johnson, The NLopt nonlinear-optimization package, http://github.com/stevengj/nlopt
https://cran.r-project.org/web/packages/nloptr/vignettes/nloptr.html
Alternatives to R
What about online tools? There are several online sites that will allow students to perform these kinds of calculations. Students familiar with MatLab know that it can be used to solve nonlinear equation problems. An open source alternative to MatLab is called GNU Octave, which can be installed on a personal computer or run online at http://octave-online.net. Students also may be aware of other sites, e.g., mycurvefit.com and IC50.tk.
Both of these free services performed well on the full dataset (results not shown), but fared poorly on the reduced subset: mycurvefit returned a value of 747.64 and IC50.tk returned an EC50 estimate of 103.6 (Michael Dohm, pers. obs.).
Simple inspection of the plotted values shows that these values are unreasonable.
EC50 calculations with Microsoft Excel
Most versions of Microsoft Excel include an add-in called Solver, which will permit mathematical modeling. The add-in is not installed as part of the default installation, but can be installed via the Options tab in the File menu for a local installation or via Insert for the Microsoft Office online version of Excel (you need a free Microsoft account). The following instructions are for a local installation of Microsoft Office 365 and were modified from information provided by sharpstatistics.co.uk.
After opening Excel, set up worksheet as follows. Note that in order to use my formulas your spreadsheet values need to be set up exactly as I describe.
- Dose values in column A, beginning with row 2
- Response values in column B, beginning with row 2
- In column F type
bin row 2,cin row 3,din row 4, andein row 5. - In cells G2 – G5, enter the starting values. For this example, I used b = 1, c = 0, d = 1, e = 400.
- For your own data you will have to explore use of different starting values.
- Enter headers in row 1.
- Column A
Dose - Column B
Response - Column C
Predicted - Column D
Squared difference - Column F
Constants
- Column A
- In cell C14 enter
sum squares - In cell D14 enter
=sum(D2:D12)
Here is an image of the worksheet (Fig. 6), with equations entered and values updated, but before Solver run.
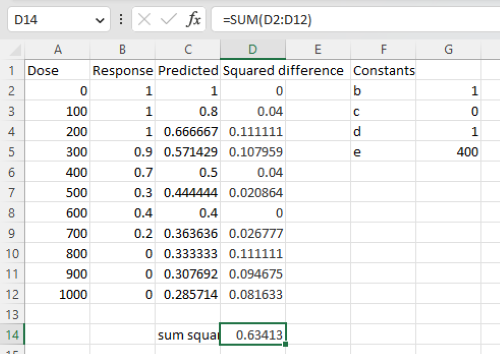
Figure 6. Screenshot Microsoft Excel worksheet containing our data set (col A & B), with formulas added and calculated. Starting values for constants in column G, rows 2 – 4.
Next, enter the functions.
- In cell C2 enter the four parameter logistic formula — type everything between the double quotes: “
=$G$3+(($G$4-$G$3)/(1+(A2/$G$5)^$G$2))“. Next, copy or drag the formula to cell C12 to complete the predictions.- Note: for a three parameter model, replace the above formula with “
=(($G$4)/(1+(A2/$G$5)^$G$2))“.
- Note: for a three parameter model, replace the above formula with “
- In cell D2 type everything between the double quotes: “
=(B2-C2)^2“. - Next, copy or drag the formula to cell
D12to complete the predictions. - Now you are ready to run solver to estimate the values for b, c, d, and e.
- Reminder: starting values must be in cells
G2:G5
- Reminder: starting values must be in cells
- Select cell
D14and start solver by clicking on Data and looking to the right in the ribbon.
Solver is not normally installed as part of the default installation of Microsoft Excel 365. If the add-in has been installed you will see Solver in the Analyze box (Fig. 7).
Figure 7. Screenshot Microsoft Excel, Solver add-in available.
If Solver is not installed, go to File → Options → Add-ins (Fig. 8).
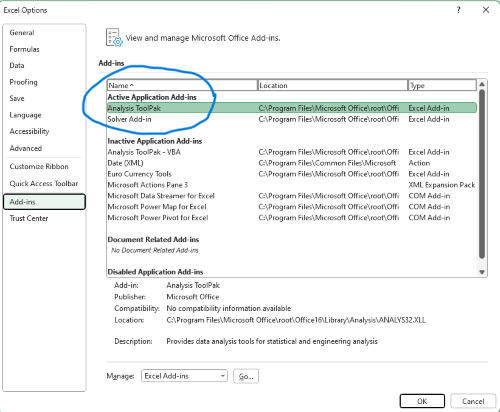
Figure 8. Screenshot Microsoft Excel, Solver add-in available and ready for use.
Go to Microsoft support for assistance with add-ins.
With the spreadsheet completed and Solver available, click on Solver in the ribbon (Fig. 7) to begin. The screen shot from the first screen of Solver is shown below (Fig. 9).
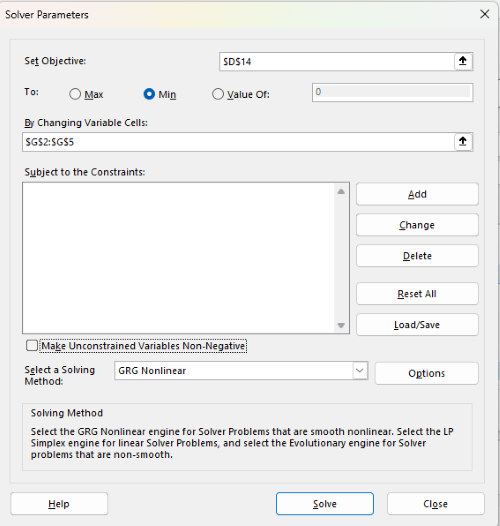
Figure 9. Screenshot Microsoft Excel solver menu.
Setup Solver
- Set Objective, enter the absolute cell reference to the sum squares value, \
$D$14 - Set To: Min.
- For By Changing Variable Cells, enter the range of cells for the four parameters, $G$2:$G$5
- Uncheck the box by “Make Unconstrained Variables Non-Negative.”
- Where constraints refers to any system of equalities or inequalities equations imposed on the algorithm.
- Select a Solving method, choose “GRG Nonlinear,” the nonlinear programming solver option (not shown in Fig. 7, select by clicking on the down arrow).
- Click Solve button to proceed.
GRG Nonlinear is one of many optimization methods. In this case we calculate the minimum of the sum of squares — the difference between observed and predicted values from the logistic equation — given the range of observed values. GRG stands for Generalized Reduced Gradient and finds the local optima — in this case the minimum or valley — without any imposed constraints. See solver.com for additional discussion of this algorithm.
If all goes well, this next screen (Fig. 10) will appear which shows the message, “Solver has converged to the current solution. All constraints are satisfied.”
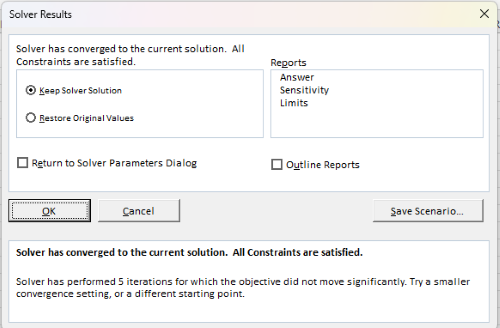
Figure 10. Screenshot solver completed run.
Click OK and note that the values of the parameters have been updated (Table 1).
Table 1. Four parameter logistic model, results from Solver
| Constant | Starting values | Values after solver |
| b | 1 | 3.723008382 |
| c | 0 | -0.088865487 |
| d | 1 | 1.017964505 |
| e | 400 | 493.9703594 |
Note the values obtained by Solver are virtually identical to the values obtained in the drc R package. The differences are probably because of the solver algorithm.
More interestingly, how did Solver do on the subset data set? Here are the results from a three parameter logistic model (Table 2).
Table 2. Three parameter logistic model, results from Solver
| Constant | Starting values | Full dataset, Solver results |
Subset, Solver results |
| b | 1 | 3.723008382 | 6.989855948 |
| d | 1 | 1.017964505 | 0.993391264 |
| e | 400 | 493.9703594 | 446.8819282 |
The results are again very close to results from the drc R package.
Thus, we would conclude that Solver and Microsoft Excel would be a reasonable choice for EC50 calculations, and much better than IC50.tk and mycurvefit.com. The advantages of R over Microsoft Excel is that the model building is more straightforward than entering formulas in the cell reference format required by Excel.
Questions
[pending]
References and suggested readings
Beck B, Chen YF, Dere W, et al. Assay Operations for SAR Support. 2012 May 1 [Updated 2012 Oct 1]. In: Sittampalam GS, Coussens NP, Nelson H, et al., editors. Assay Guidance Manual [Internet]. Bethesda (MD): Eli Lilly & Company and the National Center for Advancing Translational Sciences; 2004-. Available from: www.ncbi.nlm.nih.gov/books/NBK91994/
Di Veroli G. Y., Fornari C., Goldlust I., Mills G., Koh S. B., Bramhall J. L., Richards, F. M., Jodrell D. I. (2015) An automated fitting procedure and software for dose-response curves with multiphasic features. Scientific Reports 5: 14701.
Ritz, C., Baty, F., Streibig, J. C., Gerhard, D. (2015) Dose-Response Analysis Using R PLOS ONE, 10(12), e0146021
Tjørve, K. M., & Tjørve, E. (2017). The use of Gompertz models in growth analyses, and new Gompertz-model approach: An addition to the Unified-Richards family. PloS one, 12(6), e0178691.
Chapter 20 contents
- Additional topics
- Area under the curve
- Peak detection
- Baseline correction
- Surveys
- Time series
- Dimensional analysis
- Estimating population size
- Diversity indexes
- Survival analysis
- Growth equations and dose response calculations
- Plot a Newick tree
- Phylogenetically independent contrasts
- How to get the distances from a distance tree
- Binary classification
- Meta-analysis
/MD
20.9 – Survival analysis
draft
Introduction
As the name suggests, survival analysis is a branch of statistics used to account for death of organisms, or more generally, failure in a system. In general, this kind of analysis models time to event, where event would be death or failure. The basics of the method is defined by the survival function
![]()
where t is time, T is a variable that represents time of death or other end point, and Pr is probability of an event occurring later than at time t.
Excellent resources available, series of articles in volume 89 of British Journal of Cancer: Clark et al (2003a), Bradburn et al (2003a), Bradburn et al (2003b), and Clark et al (2003b).
Hazard function
Defined as the event rate at time t based on survival for time times equal to or greater than t.
Censoring
Censoring is a a missing data problem typical of survival analysis. Distinguish right-censored and left-censored.
Kaplan-Meier plot
Kaplan-Meier (KM) estimator of survival function. Other survival function estimators Fleming-Harrington. The KM estimator, ![]() is
is
![]()
where di is the number of events that occurred at time ti, ni is the number of individuals known to have survived or not been censored. Because it’s an estimate, a statistic, need an estimate of the error variance. Several options, the default in R is the Greenwood estimator.
![]()
The KM plot, censoring times noted with plus.
R code
Download and install the RcmdrPlugin.survival package.
Example
data(heart, package="survival")
attach(heart)
#Get help with the data set
help("heart", package="survival")
head(heart)
start stop event age year surgery transplant id 1 0 50 1 -17.155373 0.1232033 0 0 1 2 0 6 1 3.835729 0.2546201 0 0 2 3 0 1 0 6.297057 0.2655715 0 0 3 4 1 16 1 6.297057 0.2655715 0 1 3 5 0 36 0 -7.737166 0.4900753 0 0 4 6 36 39 1 -7.737166 0.4900753 0 1 4
Run basic survival analysis. After installing the RcmdrPlugin.survival, from Rcmdr select estimate survival function.
Figure 1. Screenshot of menu call for survival analysis in Rcmdr
Get survival estimator and KM plot (Figure 2)
R output
.Survfit <- survfit(Surv(start, event) ~ 1, conf.type="log", conf.int=0.95, Rcmdr+ type="kaplan-meier", error="greenwood", data=heart) .Survfit Call: survfit(formula = Surv(start, event) ~ 1, data = heart, error = "greenwood", conf.type = "log", conf.int = 0.95, type = "kaplan-meier") n events median 0.95LCL 0.95UCL 172 75 26 17 37 plot(.Survfit, mark.time=TRUE) quantile(.Survfit, quantiles=c(.25,.5,.75)) $quantile 25 50 75 3 26 67 $lower 25 50 75 1 17 46 $upper 25 50 75 12 37 NA #by default, Rcmdr removes the object remove(.Survfit)
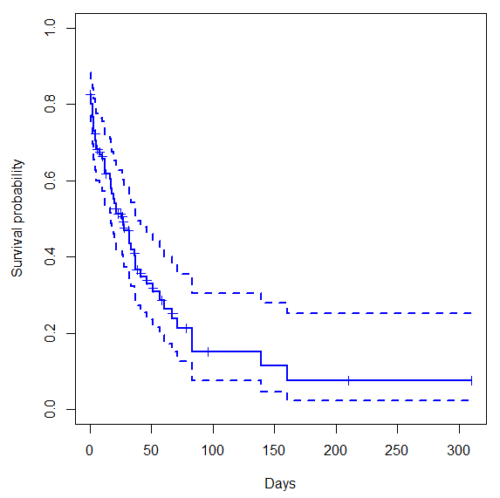
Figure 2. Kaplan-Meier plot of heart data. Dashed lines are upper and lower confidence intervals about the survival function.
Note: I modified the plot() code with these additions
ylim = c(0,1), ylab="Survival probability", xlab="Days", lwd=2, col="blue"
the data set includes age and whether or not subjects had heart surgery before transplant. Compare.
Variable surgery is recorded 0,1, so need to create a factor
fSurgery <- as.factor(surgery)
Now, to compare
Rcmdr: Statistics → Survival analysis → Compare survival functions…
R output
Rcmdr> survdiff(Surv(start,event) ~ fSurgery, rho=0, data=heart)
Call:
survdiff(formula = Surv(start, event) ~ fSurgery, data = heart,
rho = 0)
N Observed Expected (O-E)^2/E (O-E)^2/V
fSurgery=No 143 66 58.7 0.902 4.56
fSurgery=Yes 29 9 16.3 3.255 4.56
Chisq= 4.6 on 1 degrees of freedom, p= 0.03
Get the KM estimator and make a KM plot
mySurvfit <- survfit(Surv(start, event) ~ surgery, conf.type="log", conf.int=0.95,
type="kaplan-meier", error="greenwood", data=heart)
plot(mySurvfit, mark.time=TRUE, ylim=c(0,1),lwd=2, col=c("blue","red"), xlab="Number of days", ylab="Survival probability")
legend("topright", legend = paste(c("Surgery - No", "Surgery - Yes")), col = c("blue", "red"), pch = 19, bty = "n")
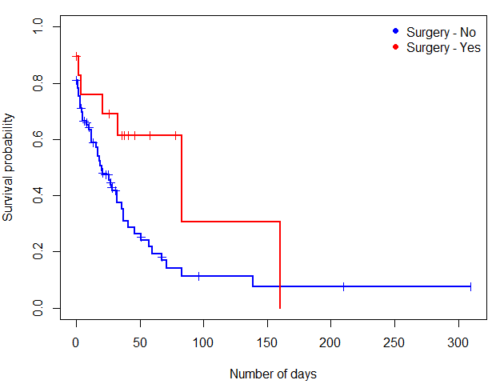
Figure 3. KM plot
The comparison plot can be made in Rcmdr by selecting our Surgery factor in Strata setting (Fig. 4). Recall that strata refers to subgroups of a population.
Figure 4. Screenshot of Survival estimator menu in Rcmdr.
Questions
[pending]
Chapter 20 contents
- Additional topics
- Area under the curve
- Peak detection
- Baseline correction
- Surveys
- Time series
- Dimensional analysis
- Estimating population size
- Diversity indexes
- Survival analysis
- Growth equations and dose response calculations
- Plot a Newick tree
- Phylogenetically independent contrasts
- How to get the distances from a distance tree
- Binary classification
- Meta-analysis
/MD
19 – Distribution free statistical methods
Introduction
We introduced the concept of permutation tests in our chapter on parameter estimates and statistical error (Chapter 3.3). Jackknife and bootstrapping are permutation approaches to working with data when the Central Limit theorem is unlikely to apply or, rather, we don’t wish to make that assumption. The jackknife is a sampling method involving repeatedly sampling from the original data set, but each time leaving one value out. The estimator, for example, the sample mean, is calculated for each sample. The repeated estimates from the jackknife approach yield many estimates which, collected, are used to calculate the sample variance. Jackknife estimators tend to be less biased than those from classical asymptotic statistics. Bootstrapping, and not jackknife resampling, is now the preferred permutation approach (add citations).
Bootstrapping
Bootstrapping involves large numbers of permutations of the data, which, in short, means we repeatedly take many samples of our data and recalculate our statistics on these sets of sampled data. We obtain statistical significance by comparing our result from the original data against how often results from our permutations on the resampled data sets exceed the originally observed results. By permutation methods, the goal is to avoid the assumptions made by large-sample statistical inference. Since its introduction, “bootstrapping” has been shown to be superior in many cases for statistics of error compared to the standard, classical approach (add citations).
Permutation vs classical NHST approach
There are many advocates for this approach, and, because we have computers now instead of the hand calculators our statistics ancestors used, permutation methods may be the approach you will take in your own work. However, the classical approach has it’s strengths; when the conditions, that is, when the assumptions of asymptotic statistics are met by the data, then the classical approaches tend to be less conservative than the permutation methods. By conservative, statisticians mean that a test performs at the level we expect it to. Thus, if the assumptions of classical statistics are met they return the correct answer more often than do the permutation tests.
Questions
[pending]
Chapter 19 contents
18 – Multiple Linear Regression
Introduction
This is the second part of our discussion about general linear models. In this chapter we extend linear regression from one (Chapter 17) to many predictor variables. We also introduce logistic regression, which uses logistic function to model binary outcome variables. Extensions to address ordinal outcome variables are also presented. We conclude with a discussion of model selection, which applies to models with two or more predictor variables.
For linear regression models with multiple predictors, in addition to our LINE assumptions, we add the assumption of no multicollinearity. That is, we assume our predictor variables are not themselves correlated.
Rcmdr and R have multiple ways to analyze linear regression models; we will continue to emphasize the general linear model approach, which allow us to handle continuous and categorical predictor variables.
Practical aspects of model diagnostics were presented in Chapter 17; these rules apply for multiple predictor variable models. Regression and correlation (Chapter 16) both test linear hypotheses: we state that the relationship between two variables is linear (the alternative hypothesis) or it is not (the null hypothesis). The difference? Correlation is a test of association (are variables correlated, we ask?), but are not tests of causation: we do not imply that one variable causes another to vary, even if the correlation between the two variables is large and positive, for example. Correlations are used in statistics on data sets not collected from explicit experimental designs incorporated to test specific hypotheses of cause and effect. Regression is to cause and effect as correlation is to association. Regression, ANOVA, and other general linear models are designed to permit the statistician to control for the effects of confounding variables provided the causal variables themselves are uncorrelated.
Models
Chapter 17 covered the simple linear model
![]()
Chapter 18 covers multiple regression linear model
![]()
where α or β0 represent the Y-intercept and β or β1, β2, … βn represent the regression slopes.
And the logistic regression model
![]()
where L refers to the upper or maximum value of the curve, ![]() refers to the rate of change at the steepest part of the curve, and
refers to the rate of change at the steepest part of the curve, and ![]() refers to the inflection point of the curve. Logistic functions are S-shaped, and typical use involves looking at population growth rates (eg, Fig 1), or in the case of logistic regression, how a treatment effects the rate of growth.
refers to the inflection point of the curve. Logistic functions are S-shaped, and typical use involves looking at population growth rates (eg, Fig 1), or in the case of logistic regression, how a treatment effects the rate of growth.
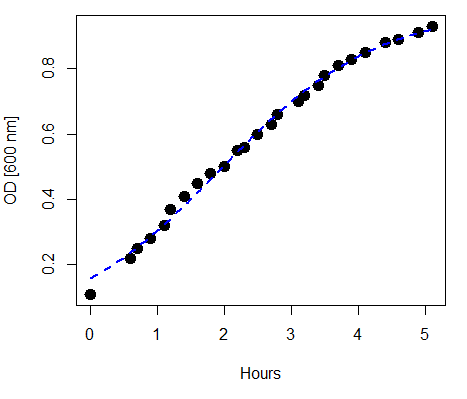
Figure 1. Growth of bacteria over time (optical density at 600 nm UV spectrophotometer), fit by logistic function (dashed line).
Suggested homework
Homework 12: Multiple linear regression.
Quizzes in this chapter
A total of 59 questions among the several subchapters, a mix of true or false and multiple choice question format.
Chapter 18 contents
17 – Linear Regression
Introduction
Regression is a toolkit for developing models of cause and effect between one ratio scale data type dependent response variables, and one (simple linear regression) or more or more (multiple linear regression) ratio scale data type independent predictor variables. By convention the dependent variable(s) is denoted by Y, the independent variable(s) represented by X1, X2, ![]() Xn for n independent variables. Like ANOVA, linear regression is simply a special case of the general linear model, first introduced in Chapter 12.7.
Xn for n independent variables. Like ANOVA, linear regression is simply a special case of the general linear model, first introduced in Chapter 12.7.
Components of a statistical model
Regression statistical methods return model estimates of the intercept and slope coefficients, plus statistics of regression fit (eg, R2, aka “R-squared,” the coefficient of determination).
Chapter 17.1 – 17.9 cover the simple linear model
![]()
Chapter 18.1 – 18.5 cover the multiple regression linear model
![]()
where α or β0 represent the Y-intercept and β or β1, β2, … βn represent the regression slopes.
Regression and correlation test linear hypotheses
We state that the relationship between two variables is linear (the alternative hypothesis) or it is not (the null hypothesis). The difference? Correlation is a test of linear association (are variables correlated, we ask?), imply possible causation, but are not sufficient evidence for causation: we do not imply that one variable causes another to vary, even if the correlation between the two variables is large and positive, for example. Correlations are used in statistics on data sets not collected from explicit experimental designs incorporated to test specific hypotheses of cause and effect.
Linear regression, however, is to cause and effect as correlation is to association. With regression and ANOVA, we are indeed making a case for a particular understanding of the cause of variation in a response variable: modeling cause and effect is the goal. Regression, ANOVA, and other general linear models are designed to permit the statistician to control for the effects of confounding variables provided the causal variables themselves are uncorrelated.
When to use correlation and when to apply linear regression to data set? For two ratio scale variables, one can always apply either approach. Correlation is always appropriate if both variables are measured in the course of the study whereas a regression modeling approach would be implied where one variable was the result of manipulation by the researcher.
Assumptions of linear regression
The key assumption in linear regression is that a straight line indeed is the best fit of the relationship between dependent and independent variables. The additional assumptions of parametric tests (Chapter 13) also hold. In Chapter 18 we conclude with an extension of regression from one to many predictor variables and the special and important topic of correlated predictor variables or multicollinearity.
Build a statistical model, make predictions
In our exploration of linear regression we begin with simple linear regression, also called ordinary least squares regression, starting with one predictor variable. Practical aspects of model diagnostics are presented. Regression may be used to describe or to provide a predictive statistical framework. In Chapter 18 we conclude with an extension of regression from one to many predictor variables. We conclude with a discussion of model selection. Throughout, use of Rcmdr and R have multiple ways to analyze linear regression models are presented; we will continue to emphasize the general linear model approach, but note that use of linear model in Rcmdr provides a number of default features that are conveniently available.
Questions
1. Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
Suggested homework
Homework 11: Simple linear regression.
Quiz
In addition to the quiz on this page, a total of 81 questions among the several subchapters, a mix of true or false and multiple choice question format.
Quiz Chapter 17
Linear Regression
References
Linear regression is a huge topic; references I include are among my favorite on the subject, but are only a small and incomplete sampling. For simplicity, I merged references for Chapter 17 and Chapter 18 into one page at References and suggested readings (Ch17 & 18)
Chapter 17 contains
- Introduction
- Simple Linear Regression
- Relationship between the slope and the correlation
- Estimation of linear regression coefficients
- OLS, RMA, and smoothing functions
- Testing regression coefficients
- ANCOVA – analysis of covariance
- Regression model fit
- Assumptions and model diagnostics for Simple Linear Regression
- References and suggested readings (Ch17 & 18)
15 – Nonparametric tests
Introduction
T-tests and ANOVA are members of a statistical family of tests called parametric tests. Parametric tests assume that the
- sample of observations come from a particular probability distribution, eg, normal distribution.
- samples among groups have equal variances.
Assumptions for parametric tests were introduced in Chapter 13. In the case of the t-test and ANOVA, we assume that the samples come from a normal probability distribution and that the probabilities of the test statistic follow the t distribution or the F distribution, respectively.
Providing these assumptions hold, we can then proceed to interpret our results as if we are talking about the population as a whole from which the samples were selected.
In other words, the t-test asks (infers) about properties of a population, hence, we are asking about parameters of the population.
T-tests, ANOVA, and other parametric tests are designed to work with quantitative ratio-scale data types. If the data are of this type and the probability distribution is known, they are the best tests to use… they allow you to make conclusions about experiments at a defined Type I error rate of 5%.
But what if you can’t assume that distribution? Your options include
- transforming the data so as the data better meet the assumptions of parametric tests.
- apply nonparametric statistical tests.
- use of distribution-free resampling methods like the bootstrap as an alternative. Bootstrapping offers methods to estimate confidence intervals and perform hypothesis tests without relying on theoretical distributions.
Here, we introduce some of the classic nonparametric statistics as an alternative to parametric tests. Bootstrapping is introduced in Chapter 19.
Nonparametric tests make fewer assumptions
Nonparametric tests do not make the assumption about a particular distribution — distribution free tests — nor are they used to make inferences about population parameters. Instead, nonparametric tests are used when the data type are ranks (ordinal). Now, when you think about it, all quantitative data can be converted to ranks. Hence, this is the argument for why there are nonparametric alternatives for tests like the t-test. There are a number of nonparametric alternatives to parametric tests and we’ll list several in this chapter (randomization methods like the bootstrap, see Ch19.2, are also considered nonparametric — unless, of course the data is assumed to come from a particular distribution, then it’s called parametric bootstrap). Another nonparametric option is to run a permutation test on the data.
However, nonparametric tests lack statistical power
One downside for nonparametric tests is that they tend to have less statistical power compared to the parametric alternatives (see Chapter 11 for review of Statistical Power). Thus, nonparametric tests tend to have higher Type II rates of error — they fail to properly reject the null hypothesis when they should. This problem tends to be less important for large sample sizes.
Note 1: Instead of transformations or other ad hoc manipulations of the data, modern statistical approaches favor modeling the error structure of the data within a Generalized Linear Model framework (St.-Pierre et al 2018). The advantage of the model approach is that parameter estimation occurs on the raw data. Use of transformations may, however, remain a better choice. While statistically justified, the generalized linear model approach may also tend to have higher rates of Type II error compared to simple transformations.
Thus, this chapter covers some of the more popular nonparametric alternative tests. Chapter 19 – Distribution free statistical methods, highlights use of permutation and randomization approaches, which also are alternatives to parametric tests.
Note 1: Bootstrapping can be either parametric or non-parametric. Non-parametric bootstrapping makes no assumptions about the data’s underlying distribution, while the parametric approach requires assuming a specific distribution.
Suggested homework
Homework 9: Common nonparametric tests.
Quizzes in this chapter
A total of 18 questions among the several subchapters, a mix of true or false and multiple choice question format.
Chapter 15 contents
14.8 – More on the linear model in Rcmdr
Introduction
During the last lectures, we could have used the Two-Way ANOVA command in R or Rcmdr
R code: How to analyze multifactorial ANOVA problems
Rcmdr: Statistics → Means → Multiway ANOVA
To analyze our two factor data sets. As long as the design meets the following conditions, by all means use this command because it is simple and precisely correct.
- Both factors are fixed, not random.
- Each level of first factor is crossed with each level of the second factor.
- No missing data (the design is fully replicated and balanced).
If any of the three points do not fit your two-way design, then you’ll need a different, more general and powerful ANOVA procedure in R and Rcmdr to analyze these types of designs. You’ll need the lm() function (Fig. 1).
Rcmdr: Statistics → Fit models → Linear model…
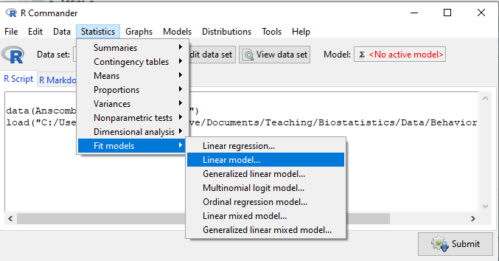
Figure 1. Linear model menu in Rcmdr, version 2.7.0
Some R basics with the lm() function, the general linear model.
![]()
Response ![]() is specified by linear predictor(s), either factors or covariates (ratio-scale predictor variables). We communicate to R what the model is by using operators. The four more commonly used operators are
is specified by linear predictor(s), either factors or covariates (ratio-scale predictor variables). We communicate to R what the model is by using operators. The four more commonly used operators are
![]() the basic way to include the model terms, i.e., the mode predictors
the basic way to include the model terms, i.e., the mode predictors
![]() which is interpreted as the interaction of all the variables and the factors in the term
which is interpreted as the interaction of all the variables and the factors in the term
![]() is interpreted as factor crossing
is interpreted as factor crossing
Not shown, but ![]() indicates the term on the left is nested within the term on the right.
indicates the term on the left is nested within the term on the right.
A few examples: we’ll have three factors, A, B, and C. For our one-way ANOVA, the model specification would be
![]()
For our crossed, balanced two-way ANOVA, the model specification would be
![]()
or equivalently
![]()
And for our block ANOVA problem?
Click here to get entire List of model commands in R.
How would we analyze our snake experiment?
Table 1. The snake data set
| Snake | Source | flick |
| 1 | dH2O | 3 |
| 2 | dH2O | 0 |
| 3 | dH2O | 0 |
| 4 | dH2O | 5 |
| 5 | dH2O | 1 |
| 6 | dH2O | 2 |
| 1 | fish | 6 |
| 2 | fish | 22 |
| 3 | fish | 12 |
| 4 | fish | 24 |
| 5 | fish | 16 |
| 6 | fish | 16 |
| 1 | worm | 6 |
| 2 | worm | 22 |
| 3 | worm | 12 |
| 4 | worm | 24 |
| 5 | worm | 16 |
| 6 | worm | 16 |
We have two factors, but one factor is a Block (repeated measures on individuals). We need to tell R and Rcmdr what our model is. We’ll return to talk about models next time, a very important topic!! For now, think of a model as adding the independent variables together to predict the response variable.
In our Snake example, it’s a two-way ANOVA, but one factor is individual snake, the other is a treatment, and we have repeat measures, so there cannot be an interaction.
We tell R and Rcmdr which columns contain the Response, and under Model, we enter the columns with the two factors.
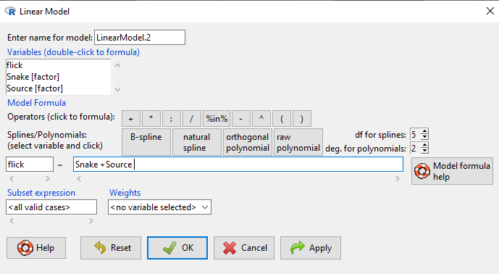
Figure 2. Menu of linear model with repeat measures model, Rcmdr, version 2.7.0.
You must also tell R and Rcmdr which factors in the model (if any) are random to get the correct F statistics. Almost without exception, blocking factors are always treated as Random
The output looks like this (see below). More complicated, true (which means more information!), but things marked in red we’ve seen before.
LinearModel.2 <- lm(flick ~ Snake +Source, data=L16SnakeTaste)
summary(LinearModel.2)
Call:
lm(formula = flick ~ Snake + Source, data = L16SnakeTaste)
Residuals:
Min 1Q Median 3Q Max
-5.2222 -0.7222 0.0278 1.5694 7.4444
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -4.444 2.523 -1.762 0.10862
Snake[T.2] 9.667 3.090 3.128 0.01072 *
Snake[T.3] 3.000 3.090 0.971 0.35451
Snake[T.4] 12.667 3.090 4.099 0.00215 **
Snake[T.5] 6.000 3.090 1.942 0.08085 .
Snake[T.6] 6.333 3.090 2.050 0.06755 .
Source[T.fish] 14.167 2.185 6.484 0.0000704 ***
Source[T.worm] 14.167 2.185 6.484 0.0000704 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 3.784 on 10 degrees of freedom
Multiple R-squared: 0.8858, Adjusted R-squared: 0.8058
F-statistic: 11.08 on 7 and 10 DF, p-value: 0.0005337
Contrast coding
When a factor (categorical variable) is used in a linear model (lm() or aov()), R must convert the factor into one or more numeric predictor variables. Contrast coding tells us how to represent the factor in the model, how to interpret the coefficients associated with the factor, how the hypothesis tests of the coefficients are handled, and whether or not Type III sums of squares tests are valid. R can handle contrast coding in a couple of ways, either as a dummy contrast (each group if the factor is contrasted to a baseline) or effects coding, where group means are compared to the grand mean. The command summary(lm()) reports contrast coding that was used in the linear model. Every kind of contrast leads to the same fitted values and same overall F-test, but coefficient estimates and t-tests change.
Note 1: Type I sums of squares tests are affected by the order of the predictors in the model, not by type of contrast coding used. R Commander defaults to Type II tests, which are generally unaffected by coding contrasts used. In contrast, Type III tests are affected by the type of coding contrasts used — Type III tests require sum-to-zero (effects) contrasts.
For the ANOVA table, we call up the command via
Rcmdr: Models → Hypothesis tests → ANOVA table… (Fig. 3).
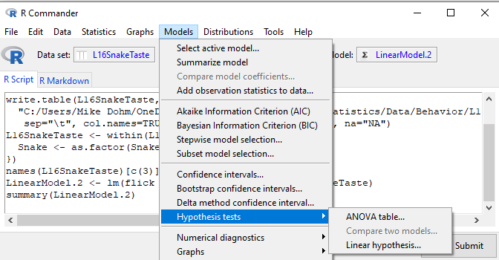
Figure 3. Rcmdr: Models → Hypothesis tests → ANOVA table… Rcmdr, version 2.7.0
Confirm that the model object is active (in this case, the object was LinearModel.2), accept the defaults about types of tests and marginality, and submit OK. The output is
Anova(LinearModel.2, type="II")
Anova Table (Type II tests)
Response: flick
Sum Sq Df F value Pr(>F)
Snake 307.61 5 4.2956 0.02396 *
Source 802.78 2 28.0256 0.00007954 ***
Residuals 143.22 10
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
End R output
You do not need to know all of the output; all of that output is there for a reason, of course, but for now, here’s what R and Rcmdr has to say (from the help menu):
The sequential sums of squares is the added sums of squares given that prior terms are in the model. These values depend upon the model order. The adjusted sums of squares are the sums of squares given that all other terms are in the model. These values do not depend upon the model order.
You should try our snake example again, but this time, remove the tongue flick response to the dH2O for the first snake — a missing value. (just type into the cell NA).
How would you use GLM to analyze a simple two-way ANOVA, a fully crossed, fully replicated (“balanced”) design? We could use the Two-way ANOVA command in R and Rcmdr, or we could use lm(). Try it with the data set from random vs nonrandom lecture.
Another example
Below, you see how the model is entered. To let R know that I wish R and Rcmdr to test the interaction, I need to add a Model term for that source of variation. I accomplish this by typing “Diet*Drug” (without the quotes).

Figure 4. Crossed, balanced design. Linear model menu, Rcmdr, version 1.9.2
After clicking OK, the following output from the lm function is returned. How does this compare to output from the two-way ANOVA command in R and Rcmdr?
You should try both and compare!
Rcmdr: Models → Hypothesis tests → ANOVA table
Anova(LinearModel.17, type="II")
Anova Table (Type II tests)
Response: chol_randomized
Sum Sq Df F value Pr(>F)
diet 141.08 2 1.3061 0.28745
drug 351.88 2 3.2577 0.05403 .
diet:drug 235.50 4 1.0901 0.38124
Residuals 1458.19 27
End R output
Nested ANOVA
The nested ANOVA may be analyzed in multiple ways in R and Rcmdr, but I prefer the lm() function because it is the most general. For Nested ANOVA, we can also use lm(). Here’s where it gets a little tricky. Put in the Response variable (Chol), then click in the box for model: Select both factors, then type in / after the factor that’s nesting factor. For our nested model example (14.5 – Nested designs), Manufacturer Source was nested within Drug.
Table 3. Nested design example data set from Chapter 14 – Nested design
| Drug | Source | Chol |
| 1 | 1 | 202.6 |
| 1 | 1 | 207.8 |
| 1 | 1 | 190.2 |
| 1 | 1 | 211.7 |
| 1 | 1 | 201.5 |
| 1 | 2 | 189.3 |
| 1 | 2 | 198.5 |
| 1 | 2 | 208.4 |
| 1 | 2 | 205.3 |
| 1 | 2 | 210 |
| 2 | 3 | 212.3 |
| 2 | 3 | 204.4 |
| 2 | 3 | 221.6 |
| 2 | 3 | 209.2 |
| 2 | 3 | 222.1 |
| 2 | 4 | 203.6 |
| 2 | 4 | 209.8 |
| 2 | 4 | 204.1 |
| 2 | 4 | 201.8 |
| 2 | 4 | 202.6 |
| 3 | 5 | 189.1 |
| 3 | 5 | 219.9 |
| 3 | 5 | 196 |
| 3 | 5 | 205.3 |
| 3 | 5 | 204 |
| 3 | 6 | 194.7 |
| 3 | 6 | 192.8 |
| 3 | 6 | 226.5 |
| 3 | 6 | 200.9 |
| 3 | 6 | 219.7 |
Note 2: If you were working with a CROSSED model, then you would enter the two factors and indicate the interaction by typing Drug*Source (if these are the two factors involved in the interaction).
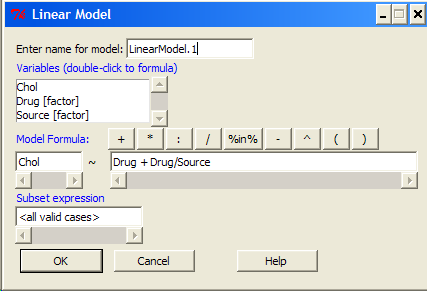
Figure 5. Nested design, linear model menu, Rcmdr, version 1.9.2
Fortunately, R and Rcmdr’s help system is quite extensive here, so when in doubt, check the help box…
Output from the linear model for the Nested Example looks like the one below.
The General Linear Model function in R and therefore Rcmdr returns information about our design plus Sums of Squares, Mean squares, and P-values. R and Rcmdr default’s to use of sequential evaluation of effects. Adjusted evaluation is useful for when you have a covariate (like body size or another confounding variable) that should be evaluated first before the factors are evaluated. We will use the sequential analysis.
Rcmdr: Models → Hypothesis tests → ANOVA table
Anova(LinearModel.15, type="II")
Anova Table (Type II tests)
Response: Chol
Sum Sq Df F value Pr(>F)
Drug 225.14 2 1.1743 0.3262
Drug:Source 269.27 3 0.9363 0.4385
Residuals 2300.61 24
End R output
Repeatability and ANOVA
We need to tell Rcmdr how to structure the error term; you need the data frame to be arranged so
aovRes <- aov(dH2O ~ Source + Error(Source/Subject), data=SnakeTaste) #Print the results aovRes Anova Table (Type II tests) Response: dH2O Sum Sq Df F value Pr(>F) Subject 307.61 5 4.2956 0.02396 * Source 802.78 2 28.0256 0.00007954 *** Residuals 143.22 10 --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 #What components are available in the aovRes object? names(aovRes) [1] "Sum Sq" "Df" "F value" "Pr(>F)"
How do I extract the “F value” for Subjects?
str(aovRes)
Questions
1. What are three learning objectives from this page?
2. Be able to distinguish use of lm() and aov() and the summary output with respect to testing of linear models.
Quiz Chapter 14.8
More on the linear model in Rcmdr
Data sets
Table 1. The snake data set
| Snake | Source | flick |
| 1 | dH2O | 3 |
| 2 | dH2O | 0 |
| 3 | dH2O | 0 |
| 4 | dH2O | 5 |
| 5 | dH2O | 1 |
| 6 | dH2O | 2 |
| 1 | fish | 6 |
| 2 | fish | 22 |
| 3 | fish | 12 |
| 4 | fish | 24 |
| 5 | fish | 16 |
| 6 | fish | 16 |
| 1 | worm | 6 |
| 2 | worm | 22 |
| 3 | worm | 12 |
| 4 | worm | 24 |
| 5 | worm | 16 |
| 6 | worm | 16 |