14.6 – Some other ANOVA designs
Introduction
There are several additional ANOVA models in common use. The crossed, balanced design is but one example of the two-way ANOVA. And, from a consideration of two factors it logically follows that there can be more than two factors as part of the design of an experiment. As the number of factors increase, the number of two-way, three-way, and even higher-order interactions are possible and at least in principle may be estimated.
Our purpose here is to highlight several, but certainly not all possible experimental designs from the perspective of ANOVA. Examples are provided. Keep in mind that the general linear model approach unifies these designs.
Some of the classical experimental ANOVA designs one sees include
- Two-way randomized complete block design
- Two-way factorial with no replication design
- Repeat-measures ANOVA with one factor
- Nested ANOVA
- Three-way ANOVA
- Split-plot ANOVA
- Latin squares ANOVA
Put simply, these designs differ in how the groups are arranged and how members of the groups are included.
Two-way randomized complete block design
This design refers to the “textbook” design. For each, factor A and factor B, there are multiple levels, in this example three levels of Factor A and three levels of Factor B, and subjects (sampling units) are randomly assigned to each level. However, one of the factors is, perhaps, of less interest, yet certainly accounts for variation in the response variable.
| Factor B | ||||
| 1 | 2 | 3 | ||
| Factor A | 1 | |||
| 2 | ||||
| 3 | ||||
where n1,1, n2,1, etc. represents the number of subjects in each cell. Thus, in this design there are nine groups. Typically, minimum replication would be three subjects per group.
Two-way factorial with no replication design
While it may seem obvious that a good experiment should have replication, there are situations in which replication is impossible. While this seems rather odd, this scenario very much describes a typical microarray, gene expression project.
| Factor B | ||||
| 1 | 2 | 3 | ||
| Factor A | 1 | |||
| 2 | ||||
| 3 | ||||
where, again, n1,1, n2,1, etc., represents the number of subjects in each cell and there are nine groups in this study. With no replication, then no more than one subject per group.
Repeated-measures ANOVA
When subjects in the study are measured multiple times for the dependent variable, this is called a repeated-measures design. We introduced the design for the simple case of before and after measures on the same individuals in Chapter 12.3. It’s straight-forward to extend the design concept to more than two measures on the subjects. The blocking effect is the individual (see Chapter 14.4), and, therefore, a random effect (see Chapters 12.3 and 14.3) in this type of experimental design.
Although straightforward in concept, repeated measure designs have many complications in practice. For example, long-term studies can expect for subjects to drop out of the study, resulting in censored data. Another complication, the assumption is that there is no carry over effect — it doesn’t matter the order different treatments are applied to the subjects. Think of this assumption as akin to the equal variances assumption in ANOVA; just like unequal variances effects Type I error rates in ANOVA, deviations from sphericity inflate Type I error rates in repeated-measures designs.
Sphericity assumption is described in two ways:
Assumption of sphericity — the ranking of individuals remains the same across treatment levels — no interaction between individual and treatment. Sphericity assumption is always met if there are just two levels of the repeated measure, e.g., before and after.
Compound symmetry assumption — the variances and covariances are equal across the study: the changes experienced by the subjects are the same across the study regardless of the order of treatments.
Tests for sphericity include:
Mauchly test: mauchly.test(object)
If results of tests for violations of sphericity warrant, corrections are available. One recommended correction is called Greenhouse-Giesser correction, which adjusts the degrees of freedom and so results in a better p-value estimate. A second correction is called Huyhn-Feldt correction; this correction, too, adjusts the degrees of freedom to improve the p-value estimate.
Three-way ANOVA
It is relatively straightforward to imagine an experiment that involves three or more factors. The analysis and interpretation of such designs, while feasible, becomes somewhat complicated especially for the mixed models (Model III).
Consider just the case of a fixed-effects 3-way ANOVA. How many tests of null hypotheses are there?
- Three tests for main effects.
- Three tests of two-way interactions.
- A test for a three-way interaction.
Thus, there are seven separate null hypotheses from a three-way ANOVA with fixed effects! As you can imagine, large sample sizes are needed for such designs, and the “higher-order” interactions (e.g., three-way interaction) can be difficult to interpret and may lack biological significance.
ANOVA designs without random assignment to treatment levels
Latin square design
We have introduced you to several ANOVA experimental designs that employed randomization for assignment of subjects to treatment groups. The purpose of randomization is even out differences due to confounding variables. However, if we know in advance something about the direction of the influence of these confounding variables, strictly random assignment is not in fact the best design. For example, the Latin square design is common in agriculture research and is very useful for situations in which two gradients are present (e.g., soil moisture levels, soil nutrient levels).
| Dry soil ←→ Wet soil | ||||
| Soil Nutrients
low |
T1 | T4 | T3 | T2 |
| T3 | T2 | T1 | T4 | |
| T2 | T3 | T4 | T1 | |
| T4 | T1 | T2 | T3 | |
Split-Plot Design
Another design from agriculture research is especially useful to ecotoxicology research. We mentioned the repeated measures design in which individuals are measured more than once and each individual receives all levels of the treatment in a random order (cross-over design). However, this design assumes that there are no carry-over effects (see Hills and Armitage 1979; For ecology/evolution definition see O’Connor et al 2014). While this assumption may hold for many experiments, we can also imagine many more situations in which this is undoubtedly false. For example, if we wish to measure the effects of ozone and relative humidity on frog behavior, we might consider using the individual as its own control. But we also wish to compare frog behavior following ozone exposure against behavior exhibited in clean air. But we are likely to violate the carry-over assumption. If a frog receives ozone then air, the effects of ozone may inhibit activity for several days after the initial exposure, which would then influence subsequent measures. The solution to this dilemma is to use what’s called a split plot design. The design combines elements of nesting.
Consider our frog experiment. There would be three factors:
Factor 1 = Exposure (air or ozone),
Factor 2 = Saturation (dry, intermediate, wet),
Factor 3 = Individual (each frog is measured 3 times).
The design table would look like
| Exposure | |||||||
| Air | Ozone | ||||||
| Humidity | Dry | Frog1 | Frog2 | Frog3 | Frog4 | Frog5 | Frog6 |
| Intermediate | Frog1 | Frog2 | Frog3 | Frog4 | Frog5 | Frog6 | |
| Wet | Frog1 | Frog2 | Frog3 | Frog4 | Frog5 | Frog6 | |
Thus, the design is crossed for one factor (saturation), but nested for another factor (individuals are nested within Exposure factor).
Questions
- Which of the study designs mentioned so far are sensitive to carry-over effects?
- With respect to how levels of Factors are assigned, distinguish the split-plot design from the Latin square design.
Quiz Chapter 14.6
Some other ANOVA designs
Chapter 14 contents
14.5 – Nested designs
Introduction
Crossed versus nested design
Factors are independent variables whose values we control and wish to study because we believe they have an effect on the dependent variable. While it is logical to think of factors and levels within factors as independent variables fully under our control, a moments reflection will come up with examples in which the groups (levels) depend on the factor.
Crossed – each level of a factor is in each level of the other factor. This was illustrated in Chapter 14.1 on Crossed, balanced, fully replicated two-way ANOVA.
Nested – levels of one factor are NOT the same in each of the levels of the other factor. Nested designs are an important experimental design in science, and they have some advantages over the 2-way ANOVA design (for one), but they also have limitations.
Classic examples of nesting: culturing and passage of cell lines in routine cell colony maintenance means that even repeated experiments are done on different experimental units. Cells derived from one vial are different from cells derived from a different vial. Similarly, although mice from an inbred strain are thought to be genetically identical, environments vary across time, so mice from the same strain but born or purchased at different times are necessarily different. These scenarios involving time create a natural block effect. Thus, cells are nested by block effect passage number and mice are nested by block effect colony time. We introduced randomized block design in the previous Chapter 14.4.
Statistical model
If Factor B is nested within Factor A, then a group or level within Factor B occurs only within a level of Factor A. Like the randomized block model, there will be no way to estimate the interaction in a nested two-way ANOVA. Our statistical model then is
![]()
Examples
Example 1. Three different drugs, 6 different sources of the drugs. The researcher obtains three different drugs from 6 different companies and wants to know if one of the drugs is better than another drug (Factor A) in lowering the blood cholesterol in women. There is always the possibility that different companies will be better or worse at making the drug. So the researchers also use the Factor Source (Factor B) to examine this possibility. Unfortunately they can not obtain all drugs from the same sources. This leads to a Nested ANOVA — notice that each drug is obtained from a different source.
We CANNOT perform the typical two-factor ANOVA because we cannot get a mean of the different drugs by combining the same levels of the Sources: the data is NOT crossed. The Sources of the drugs (Factor B) are NESTED within the type of Drug (Factor A): each source is only found in one of the Drug categories. So, we can’t calculate a mean for the Drug levels independent of the SOURCE from which the drug came.
Table 1. Example of a nested design
|
Drug A
|
Drug B
|
Drug C
|
|||
|
Source 1
|
Source 2
|
Source 3
|
Source 4
|
Source 5
|
Source 6
|
|
202.6
|
189.3
|
212.3
|
203.6
|
189.1
|
194.7
|
|
207.8
|
198.5
|
204.4
|
209.8
|
219.9
|
192.8
|
|
190.2
|
208.4
|
221.6
|
204.1
|
196.0
|
226.5
|
|
211.7
|
205.3
|
209.2
|
201.8
|
205.3
|
200.9
|
|
201.5
|
210.0
|
222.1
|
202.6
|
204.0
|
219.7
|
Scroll to end of this page to get the data set in stacked worksheet format, or click here.
Compare Table 1 to CROSSED data structure (Table 2) — a typical two-factor ANOVA — which would look like
Table 2. Contents of Table 1 presented as crossed design
|
Drug A
|
Drug B
|
Drug C
|
|||
|
Source 1
|
Source 2
|
Source 1
|
Source 2
|
Source 1
|
Source 2
|
|
202.6
|
189.3
|
?
|
?
|
?
|
?
|
|
207.8
|
198.5
|
?
|
?
|
?
|
?
|
|
190.2
|
208.4
|
?
|
?
|
?
|
?
|
|
211.7
|
205.3
|
?
|
?
|
?
|
?
|
|
201.5
|
210.0
|
?
|
?
|
?
|
?
|
We can take a mean of the different drugs by combining the same levels of the Sources. Here’s the nested design (Table 3).
Table 3. Group means, nested design
|
Drug A
|
Drug B
|
Drug C
|
|||
|
Source 1
|
Source 2
|
Source 3
|
Source 4
|
Source 5
|
Source 6
|
|
202.76
|
202.3
|
213.92
|
204.38
|
202.86
|
206.92
|
We can take a mean of the different drugs by combining the same levels of the Sources. Here’s the crossed design (Table 4).
Table 4. Groups means by crossed design.
|
Drug A
|
Drug B
|
Drug C
|
|||
|
Source 1
|
Source 2
|
Source 1
|
Source 2
|
Source 1
|
Source 2
|
|
202.76
|
202.3
|
?
|
?
|
?
|
?
|
Why the “?” in Table 2 and 4? Manufacturing source 1 & 2 do not sell Drug B and Drug C. So, there cannot be a crossed design.
Why can’t we just use a One-Way ANOVA? Can’t we just ANALYZE the three DRUGS separately, ignoring the source issue (after all, the drugs are not all made by the same manufacturer)? But it is not a one-way ANOVA problem… Here’s why.
The researcher suspects that the response of a particular drug might be dependent upon the particular source from which the drug was purchased. So, the type of source from which the drug was purchased is another FACTOR. Thus, drugs from one source might have more (less) affect compared to drugs from another source regardless of the type of drug. However, each drug is NOT available from each source. Thus the research design can NOT be crossed and Drug is NESTED within Source.
We can ask ONLY two questions (hypotheses) from this NESTED ANOVA research design:
HO: There is no difference in the average effect of the drugs on (tumor size, cholesterol level, blood pressure, etc.)
HA: There is a difference in the average effect of the drugs on (tumor size, cholesterol level, blood pressure, etc.)
HO: There is no difference in the average effect of the drugs on (tumor size, cholesterol level, blood pressure, etc.) purchased from different manufacturers.
HA: There is a difference in the average effect of the drugs on (tumor size, cholesterol level, blood pressure, etc.) purchased from different manufacturers.
Notice that we do NOT examine the effect of the interaction between Drug type and source of the drug. Why not?
Table 5. Sources of Variation in Nested ANOVA
| Source of Variation |
Sum of Squares
|
DF
|
Mean Squares
|
| Total | |||
| Among all subgroups |
|
||
| Among Groups |
|
||
| Among Subgroups | |||
| Error |
Subtract all of the subgroup Sums of Squares from the Total Sums of Squares
|
Testing nested ANOVA with one main factor
Perhaps surprisingly given the number of terms above, there are only two hypothesis tests, and, only one of REAL interest to us. There are exceptions (e.g., quantitative genetics provides many examples), but we are generally most interested in the among group test — this is the test of the main factor. In our example, the main factor was DRUG and whether the drugs differed in their effects on cholesterol levels. The second test is important in the sense that we prefer that it contributes little or no variation to the differences in cholesterol levels. But it might.
Table 6. F statistics for nested ANOVA
| F for the main effect is given as | |
| F for the subgroup is given by | |
| and of course, use the appropriate DF when testing the F values!! The Critical Value F0.05 (2), df numerator, df denominator | |
One way to look at this: it would not make sense to conclude that an effect of the main group was significant if the variation in the subgroups was much, much larger. That’s in part why we test the main effect with the subgroups MS and not the error MS. If variation due to the nested variable is not significant, then it is an estimate of the error variance, too.
The nested model we are describing is a two factor ANOVA, but it is incomplete (compared to the balanced, fully crossed 2-way design we’ve talked about before). We don’t have scores in every cell. Instead, each level of nested factor is paired with one and only one level of the other factor. In our example, Source is paired with only one other level of the other factor Drug (e.g., Source 1 goes with Drug 1 only), but the main effect is paired with 2 levels of the nesting factor (e.g., Drug 1 is manufactured at Source 1 and Source 2).
Note that nesting is strictly one way. Drug is not nested within source, for example.
Some important points about testing the null hypotheses in a nested design. For one, the test of the effect of the nesting factor (Source) is confounded by the interaction between the main factor. We don’t actually know if the interaction is present, but we also get no way to test for it because of the incomplete design. We must therefore be cautious in our interpretation of the effect of the nested factor.
Consider our example. We want to interpret the effect of source as the contribution to the response based on variation among the different suppliers of the drugs. It might be good to know that some drug manufacturer is better (or worse) than others. However, differences among the sources for the different drugs are completely contained in the main effect factor (the test of effects of the different drugs themselves on the response). Therefore, the observed differences between sources COULD be entirely due to the effects of the different drugs and have nothing to do with variation among sources!!
Questions
- Identify the response variable and whether the described factor (in all caps) is suitable for crossed design or nested design
a. In a breeding colony of lab mice, BREEDERS are used to generate up to five LITTERS; effects on offspring REPRODUCTIVE SUCCESS.
b. Effects of individual TEACHERS at different SCHOOLS on STUDENT LEARNING in biology.
c. Lisinopril, an ACE-inhibitor drug prescribed for treatment of high blood pressure, is now a generic drug, meaning a number of COMPANIES can manufacture and distribute the medication. Millions of DOSES of lisinopril are made each year; drug companies are required by the FDA to record when a dose is made and to record these dates by LOT NUMBER. - Work the example data set provide in this page. After loading the data set into Rcmdr (R), use linear model. The command to nest requires use of the forward slash, /. For example, if factor b is nested within factor a, then a/b. The linear model formula then,
Model <- lm(Obs ~ a/b, data=source)
- Describe the problem, i.e., what is a? What is b? What are the hypotheses?
- What is the statistical model?
- Test the model.
- Conclusions?
Quiz Chapter 14.5
Nested designs
Data set used in this page
| Drug | Source | Obs |
| A | s1 | 202.6 |
| A | s1 | 207.8 |
| A | s1 | 190.2 |
| A | s1 | 211.7 |
| A | s1 | 201.5 |
| A | s2 | 189.3 |
| A | s2 | 198.5 |
| A | s2 | 208.4 |
| A | s2 | 205.3 |
| A | s2 | 210 |
| B | s3 | 212.3 |
| B | s3 | 204.4 |
| B | s3 | 221.6 |
| B | s3 | 209.2 |
| B | s3 | 222.1 |
| B | s4 | 203.6 |
| B | s4 | 209.8 |
| B | s4 | 204.1 |
| B | s4 | 201.8 |
| B | s4 | 202.6 |
| C | s5 | 189.1 |
| C | s5 | 219.9 |
| C | s5 | 196 |
| C | s5 | 205.3 |
| C | s5 | 204 |
| C | s6 | 194.7 |
| C | s6 | 192.8 |
| C | s6 | 226.5 |
| C | s6 | 200.9 |
| C | s6 | 219.7 |
Chapter 14 contents
14.4 – Randomized block design
Introduction
Randomized Block Designs and Two-Factor ANOVA
In previous lectures, we have introduced you to the standard factorial ANOVA, which may be characterized as being crossed, balanced, and replicated. We expect additional factors (covariates) may contribute to differences among our experimental units, but rather than testing them — which would increase need for additional experimental units because of increased number of groups to test — we randomize our subjects. Randomization is intended to disrupt trends of confounding variables (aka covariates). If the resulting experiment has missing values (see Chapter 5), then we can say that the design is partially replicated; if only one observation is made per group, then the design is not replicated — and perhaps, not very useful!!
A special type of Two-factor ANOVA which includes a “blocking” factor and a treatment factor.
Randomization is one way to control for “uninteresting” confounding factors. Clearly, there will be scenarios in which randomization is impossible. For example, it is impossible to randomly assign subjects to The blocking factor is similar to the 10.3 – Paired t-test. In the paired t-test we had two individuals or groups that we paired (e.g. twins). One specific design is called the Randomized Block Design and we can have more than 2 members in the group. We arrange the experimental units into similar groups, i.e., the blocking factor. Examples of blocking factors may include day (experiments are run over different days), location (experiments may be run at different locations in the laboratory), nucleic acid kits (different vendors), operator (different assistants may work on the experiments), etcetera.
In general we may not be directly interested in the blocking factor. This blocking factor is used to control some factor(s) that we suspect might affect the response variable. Importantly, this has the effect of reducing the sums of squares by an amount equal to the sums of squares for the block. If variability removed by the blocking is significant, Mean Square Error (MSE) will be smaller, meaning that the F for treatment will be bigger — meaning we have a more powerful ANOVA than if we had ignored the blocking.
Statistical Testing in Randomized Block Designs
“Blocks” is a Random Factor because we are “sampling” a few blocks out of a larger possible number of blocks. Treatment is a Fixed Factor, usually.
The statistical model is
![]()
The Sources of Variation are simpler than the more typical Two-Factor ANOVA because we do not calculate all the sources of variation – the interaction is not tested! (Table 1).
Table 1. Sources of variation for a two-way ANOVA, randomized block design
|
Sources of Variation & Sum of Squares
|
DF
|
Mean Squares
|
![]()
Critical Value F0.05(2), (a – 1), (Total DF – a – b)
In the exercise example above: Factor A = exercise or management plan.
Notice that we do not look at the interaction MS or the Blocking Factor (typically).
Learn by doing
Rather than me telling you, try on your own. We’ll begin with a worked example, then proceed to introduce you to three problems. Click here (Ch 14.8)for general discussion of Rcmdr and linear models for models other than the standard 2-way ANOVA (Chapter 14.8).
Worked example
Wheel running by mice is a standard measure of activity behavior. Even wild mice will use wheels (Meijer and Roberts 2014). For example, we conduct a study of family parity to see if offspring from the first, second, or third sets of births from wheel-running behavior in mice (total revs per 24 hr period). Each set of offspring from a female could be treated as a block. Data are for 3 female offspring from each pairing. This type of data set would look like this:
Table 2. Wheel running behavior by three offspring from each of three birth cohorts among four maternal sets (moms).
|
Revolutions of wheel per 24-hr period
|
||||
|
Block
|
Dam 1
|
Dam 2
|
Dam 3
|
Dam 4
|
|
b1
|
1100
|
1566
|
945
|
450
|
|
b1
|
1245
|
1478
|
877
|
501
|
|
b1
|
1115
|
1502
|
892
|
394
|
|
b2
|
999
|
451
|
644
|
605
|
|
b2
|
899
|
405
|
650
|
612
|
|
b2
|
745
|
344
|
605
|
700
|
|
b3
|
1245
|
702
|
1712
|
790
|
|
b3
|
1300
|
612
|
1745
|
850
|
|
b3
|
1750
|
508
|
1680
|
910
|
Thus, there were nine offspring for each female mouse (Dam1 – Dam4), three offspring per each of three litters of pups. The litters are the blocks. We need to get the data stacked to run in R. I’ve provided the full dataset for readers, scroll to end of this page or click here.
Question 1. Describe the problem and identify the treatment and blocking factors.
Answer. Each female has three litters. We’re primarily interested in genetics (and maternal environment) of wheel running behavior, which is associated with the moms (Treatment factor). The questions is whether there is an effect of birth parity on wheel running behavior. Offspring of a first-time mother may experience different environment than offspring of experienced mother. In this case, parity effects is an interesting question, nevertheless, blocking is the appropriate way to handle this type of problem.
Question 2. What is the statistical model?
Answer. Response variable, Y, is wheel running. Let α the effect of Dam and β the birth cohorts (i.e., the blocking effect).
![]()
Question 3. Test the model.
Answer. We fit the main effects, Dam and Block Fig 1.
![]()
Rcmdr: Statistics → Fit models → Linear model
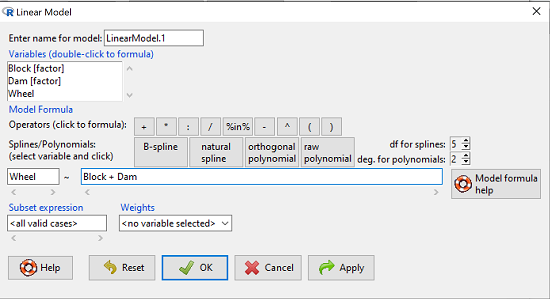
Figure 1. Screenshot Rcmdr Linear Model menu.
then run the ANOVA summary to get the ANOVA table. Rcmdr: Models → Hypothesis tests → ANOVA table.
R Output
Anova Table (Type II tests)
Response: Wheel
Sum Sq Df F value Pr(>F)
Dam 1467020 3 4.4732 0.01036 *
Block 1672166 2 7.6482 0.00207 **
Residuals 3279544 30
Question 4. Conclusions?
Answer 4. The null hypotheses are:
Treatment factor: Offspring of the different dams have same wheel running activity of offspring.
Blocking factor: No effect of litter parity on wheel running activity of offspring.
Both the treatment factor (p = 0.01036) and the blocking factor (p = 0.00207) were statistically significant.
Questions
Problem 1.
Or we might want to measure the Systolic Blood Pressure of individuals that are on different exercise regimens. However, we are not able to measure all the individuals on the same day at the same time. We suspect that time of day and the day of the year might effect an individuals blood pressure. Given this constraint, the best research design in this circumstance is to measure one individual on each exercise regime at the same time. These different individuals will then be in the same “block” because they share in common the time that their blood pressure was measured. This type of data set would look like this (Table 2):
Table 2. Simulated blood pressure of five subjects on three different exercise regimens.†
|
Subject
|
No
Exercise |
Moderate Exercise
|
Intense Exercise
|
|
1
|
120
|
115
|
114
|
|
2
|
135
|
130
|
131
|
|
3
|
115
|
110
|
109
|
|
4
|
112
|
107
|
106
|
|
5
|
108
|
103
|
102
|
Let’s make a line graph to help us visualize trends (Fig 2).
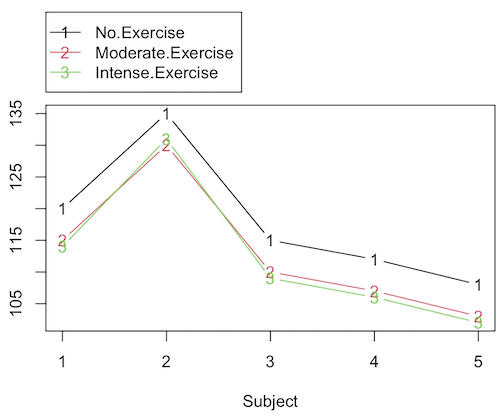
Figure 2. Line graph of data presented in Table 2.
Question 1. Describe the problem and identify the treatment and blocking factors.
Question 2. What is the statistical model?
Question 3. Test the model.
Question 4. Conclusions?
†You’ll need to arrange the data like the data set for the worked example.
Problem 2.
Another example in conservation biology or agriculture. There may be three different management strategies for promoting the recovery of a plant species. A good research design would be to choose many plots of land (blocks) and perform each treatment (management strategy) on a portion of each plot of land (block). A researcher would start with an equal number of plantings in the plots and see how many grew. The plots of land (blocks) share in common many other aspects of that particular plot of land that may effect the recovery of a species.
Table 3. Growth of plants in 5 different plots subjected to one of three management plans (simulated data set).†
|
Plot No.
|
No Management Used
|
Management
Plan 1 |
Management
Plan 2 |
|
1
|
0
|
11
|
14
|
|
2
|
2
|
13
|
15
|
|
3
|
3
|
11
|
19
|
|
4
|
4
|
10
|
16
|
|
5
|
5
|
15
|
12
|
†You’ll need to arrange the data the same arrangement as for the worked example.
These are examples of Two-Factor ANOVA but we are usually only interested in the treatment Factor. We recognize that the blocking factor may contribute to differences among groups and so wish to control for the fact that we carried out the experiments at different times (e.g., seasons) or at different locations (e.g., agriculture plots)
Question 5. Describe the problem and identify the treatment and blocking factors. Make a line graph to help visualize.
Question 6. What is the statistical model?
Question 7. Test the model.
Question 8. Conclusions?
Repeated-Measures Experimental Design
If multiple measures are taken on the same individual, then we have a repeated-measures experiment. This is a Randomized Block Design. In other words, each animal gets all levels of a treatment (assigned randomly). Thus, samples (individuals) are not independent and the analysis needs to take this into account. Just like for paired-T tests, one can imagine a number of experiments in biomedicine that would conform to this design.
Problem 3.
The data are total blood cholesterol levels for 7 individuals given 3 different drugs (from example given in Zar 1999, Ex 12.5, pp. 257-258).
Table 5. Repeated measures of blood cholesterol levels of seven subjects on three different drug regimens.†
|
Subjects
|
Drug1
|
Drug2
|
Drug3
|
|
1
|
164
|
152
|
178
|
|
2
|
202
|
181
|
222
|
|
3
|
143
|
136
|
132
|
|
4
|
210
|
194
|
216
|
|
5
|
228
|
219
|
245
|
|
6
|
173
|
159
|
182
|
|
7
|
161
|
157
|
165
|
†You’ll need to arrange the data like the data set for the worked example.
Question 9: Is there an interaction term in this design? Make a line graph to help visualize.
Question 10: Are individuals a fixed or a random effect?
Question 11. What is the statistical model?
Question 12. Test the model. Note that we could have done the experiment with 21 randomly selected subjects and a one-factor ANOVA. However, the repeated measures design is best IF there is some association (“correlation”) between the data in each row. The computations are identical to the randomized block analysis.
Question 13. Conclusions?
Problem 4.

Figure 3. Juvenile garter snake, image from juvenile garter snake in hand, public domain.
Here is a second example of a repeated measures design experiment. Garter snakes (Fig 3) respond to odor cues to find prey. Snakes use their tongues to “taste” the air for chemicals, and flick their tongues rapidly when in contact with suitable prey items, less frequently for items not suitable for prey. In the laboratory, researchers can test how individual snakes respond to different chemical cues by presenting each snake with a swab containing a particular chemical. The researcher then counts how many times the snake flicks its tongue in a certain time period (data presented p. 301, Glover and Mitchell 2016).
Table 6. Tongue flick counts of naïve newborn snakes to extracts†
|
Snake
|
Control (dH2O)
|
Fish mucus
|
Worm mucus
|
|
1
|
3
|
6
|
6
|
|
2
|
0
|
22
|
22
|
|
3
|
0
|
12
|
12
|
|
4
|
5
|
24
|
24
|
|
5
|
1
|
16
|
16
|
|
6
|
2
|
16
|
16
|
†You’ll need to arrange the data like the data set for the worked example.
Question 14. Describe the problem and identify the treatment and blocking factors. Make a line graph to help visualize.
Question 15. What is the statistical model?
Question 16. Test the model.
Question 17. Conclusions?
Quiz Chapter 14.4
Randomized block design
Data sets used in this page
Problem 1 data set
| Block | Dam | Wheel |
|---|---|---|
| B1 | D1 | 1100 |
| B1 | D2 | 1566 |
| B1 | D3 | 945 |
| B1 | D4 | 450 |
| B1 | D1 | 1245 |
| B1 | D2 | 1478 |
| B1 | D3 | 877 |
| B1 | D4 | 501 |
| B1 | D1 | 1115 |
| B1 | D2 | 1502 |
| B1 | D3 | 892 |
| B1 | D4 | 394 |
| B2 | D1 | 999 |
| B2 | D2 | 451 |
| B2 | D3 | 644 |
| B2 | D4 | 605 |
| B2 | D1 | 899 |
| B2 | D2 | 405 |
| B2 | D3 | 650 |
| B2 | D4 | 612 |
| B2 | D1 | 745 |
| B2 | D2 | 344 |
| B2 | D3 | 605 |
| B2 | D4 | 700 |
| B3 | D1 | 1245 |
| B3 | D2 | 702 |
| B3 | D3 | 1712 |
| B3 | D4 | 790 |
| B3 | D1 | 1300 |
| B3 | D2 | 612 |
| B3 | D3 | 1745 |
| B3 | D4 | 850 |
| B3 | D1 | 1750 |
| B3 | D2 | 508 |
| B3 | D3 | 1680 |
| B3 | D4 | 910 |
Chapter 14 contents
14.3 – Fixed effects, Random effects
Introduction
With few exceptions (eg, repeatability and intraclass correlation calculations, Chapter 12.3), we have been discussing the Model I ANOVA or fixed-effects ANOVA– fixed implies that we select the levels for the factor. It may not be obvious — in hindsight it is — but levels may also be randomly selected, eg, nature provides the levels. Thus, levels are random and the model is a random-effects ANOVA. Beginning with our discussions of ANOVA, it becomes increasingly important to incorporate concept of models in statistics. As you have been working in R and Rcmdr with the lm() function, you have been forced to address the statistical model concept — you enter the response variable then type in both factors and create a term for the interaction.
We’ve just completed an experiment in which the response (cholesterol levels) of 36 individuals, from one of three drug treatments (placebo, Drug A, Drug B), given one of 3 types of diets (low, medium, or high carbohydrate), was observed. Thus, we say that the specific treatments of drug and diet contribute to variation in cholesterol levels. More formally, we say that the observed response of the kth individual (Yijk) is equal to the overall mean (m) plus the added effect of Drug (a) plus the added effect of Diet (b) plus the interaction between Diet and Population (ab) plus unidentified sources of variation generally called error (e). In symbols, we write
![]()
where i is the number of levels of the first factor (in our example, Diet had 2 levels, so i = A or i = B), j is the number of levels of the second factor (in our example, Population had 2 levels, so j = 1, or j = 2), and k is the total number of observations in the experiment (12 in our example, so k = 1, 2, …, 11, 12). Thus, we can think of each term “adding up” to give us the observed value.
Although it is a bit confusing at first, these equations help us understand how the experiment was conducted and therefore how to analyze and interpret the results.
Model I, Model II & Model III ANOVA
Statisticians recognize that how levels of the factors were selected for an experiment impact conclusions from ANOVAs. The key: whether or not the levels of the factor were selected (1) randomly from all possible levels of the factor or (2) specifically selected by the experimenters. We introduced the concepts of fixed and random effects in Chapter 12.3. For one-way ANOVA, the distinction between fixed and random effects influences the interpretation, but not the calculation of the ANOVA components. For two or more treatment factors, both the interpretations and the calculations of ANOVA components are affected.
There are two general types of Factors that we can choose to employ in an ANOVA: Fixed Model ANOVA and Random Model ANOVA. Where two or more factors apply, by far the most common model in experimental sciences is a combination of fixed and random, so we need to add a third general type, the Mixed Model ANOVA design.
Fixed Factors. Where the levels of the factor are selected by us. In this case we would only be interested in the response of the individuals that are given those specific treatments.
Medicine – for example, where we choose a treatment given to patients with a history of coronary heart disease; compare outcomes of patients given a statin (drug used to lower serum cholesterol) drug versus a placebo. (Note that this is the same study we discussed in our lecture on about risk analysis).
Ecotoxicology – for example, compare growth rates and deformities of tadpole frogs given Aldicarb, Atrazine (both estrogen-mimicking pesticides), or a control (ie, no pesticides). If you’re interested in these topics, here’s a link to the EPA’s web site, listing pesticide sales and use in the United States. Here’s a link to a NIH National Institute of Environmental Sciences, with a nice description of estrogen mimicking pesticides.
Agriculture & Genetics – for example, monitor growth of a particular hybrid corn available from three different manufacturers. See an example of Wisconsin corn hybrid studies.
In each of these examples we might be interested in those specific treatments and no other treatments.
HO: No difference in the means among the levels of the Factor
HA: Some difference in the means among these specific levels of the Factor; the specific levels of this factor affects the response variable.
This is an example of a Model I ANOVA, also called a “fixed effects” model ANOVA.
Random Factors. We still only use a relatively few number of different levels of a particular factor. However, in this case we are interested in many different levels of the factor — we want to generalize beyond our sample. The levels that we use would be a “sample” of all possible levels that we would be interested in.
Medicine – for example, where we randomly choose a treatment level given to patients; four concentrations of a drug and a placebo. Since concentration is a ratio scale data type, concentration can range from 0 (the placebo) to 100%.
Ecotoxicology – for example, release different concentrations or mixtures of air plus components of air pollution to chambers, record the response of plants or animals.
Agriculture & Genetics – for example, grow three different varieties of a plant in three different soils or different genetic strains of animals on three different diets. In these cases, factor levels are random because we are drawing from a large pool of possible levels: genetic varieties or strains — we selected three, but it’s rare that were are specifically interested in the three chosen. More often, we want to make generalizations and the three were somehow representative (we hope) of genetic variation in the species of interest.
In each of these examples, we write the null hypothesis to reflect that the particular levels are only of interest in so far as they can be used to generalize back to the population.
HO: No difference in the variation between groups.
HA: Some difference in the variation among these groups.
This is an example of a Model II ANOVA, also called a “random effects” model ANOVA. Note that we specify the hypothesis in terms of variation, not of the means.
Your two-way ANOVA could be Model I, Model II, or it could be mixed, with one factor fixed, the other random (this later model is called a Model III, or “mixed model” ANOVA).
For the most part, the distinction between whether you have fixed or random effects is clear, but whether we use fixed or random or combinations, this design decision has consequences for testing.
The decision does not affect the Sources of Variation for the different Models. Last time, we showed the tests for a Model I ANOVA (the “fixed effects model”).
For Random Effects or Mixed Effects, we only change how we determine the statistical significance of the Factors. Here’s a summary of how the experimental design changes the calculation of the F-test statistic for the Factors.
Table 1. Calculation of F
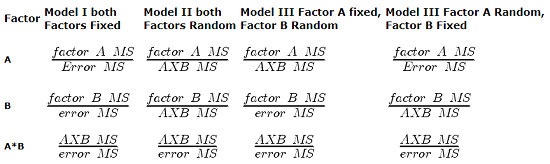

The Critical Value for each of the different F values will be obtained by simply finding the degrees of freedom for the numerator and denominator SS. This was discussed and can be found sources of variation in 2-way ANOVA lecture.
From the formulas, we can see that the major difference is that sometimes the Factor MS is divided by the error MS and sometimes it is divided by the interaction MS.
If the interaction term is NOT statistically significant, then the Interaction MS (mean square) estimates the Error MS. In other words, if the interaction term is not statistically significant it will be similar in magnitude as the Error MS. In this case there will be no large difference in the computed outcomes if the Factor A or B is fixed or random.
However, there will be times when the interaction is not significant but the interaction MS is still larger than the Error MS. Then there could be a difference in the F value for the Factor.
If the interaction term is Significant and the interaction MS is larger than the Error MS then there will be difference in F values for the Factors A and/or B. The F values will be smaller for the Factors MS that are divided by the interaction MS.
It is possible that they will become non-significant with the interaction MS as the denominator. So it will become Harder to detect a significant Factor effect if there is also a significant Interaction effect. A graphical representation will help us understand why we use the interaction MS in some instances as the denominator.
In fact, if the interaction is found to be statistically significant, we must then interpret the effects of factors with caution. In general, if the interaction is significant, then the main factors are generally not interpreted in the 2-way ANOVA. Instead, a series of one-way ANOVA’s are conducted holding one of the factors constant. For example, evaporative water loss (EWL) in frogs in the presence of air pollution (ozone) may depend on the relative humidity (RH) — if the RH is low, the frog may lose less body water at different concentrations of ozone than if the RH is moderate. Therefore, since the interaction is significant, the best thing to do is to look at the effects of ozone concentration on EWL at each level of saturation (RH).
This is a critical point in your understanding of complex ANOVA designs. Let us examine a case where there is a mildly significant interaction effect between two factors. In the first graph below (Fig 1) we see that Genotype 2 performs better on average (combining the two density treatments). If we are only interested in these two density treatments then it might be that the Genotype Factor is significant. This would be the case if Factor A is fixed and Factor B (density) is also fixed.
The Formula for Factor A would be: F = Genotype MS / error MS
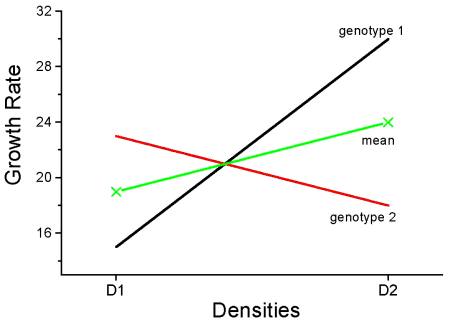
Figure 1. Interaction example. At density D1, genotype 2 (red line) has higher growth rate; at density D2, the ranking switches: now, genotype 1 (black line) has higher growth rate.
However, it is likely that both Factor A (genotype) and Factor B (densities) are actually “samples” of many other possible genotypes and densities that we could examine.
Consider more than two genotypes raised in more than 2 densities (Fig 2). The outcome might look like
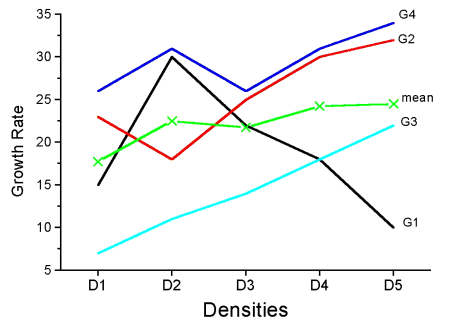
Figure 2. Interaction example expanded for multiple genotypes over multiple densities.
The graph (Fig 2) shows that genotype 2 does not do better than genotype 1 if we have more densities. If we also have other genotypes we see that there are other genotypes that have better (higher) responses than genotype 2.
![]()
In these cases it would have been more appropriate to calculate the F value for Factor A (genotype) using the interaction MS as the denominator. In Figure 1 there was some interaction this will make it harder to reject the null hypothesis that there is no effect of Factor A (genotype). So you must be careful to think about how you plan to interpret your data before you decide how to analyze the data using a Two-Factor ANOVA.
Questions
- Which of the following statements regarding fixed and random factors is true?
A. With fixed factors, the subjects are selected by the researcher
B. With fixed factors, the treatment levels are selected by the researcher at random from all possible levels
C. With fixed factors, the subjects are selected at random by the researcher
D. With random factors, the treatment levels are selected by the researcher at random from all possible levels - Please write the equation for the one-way ANOVA with four levels of of fixed effects treatment factor A (you may wish to review Chapter 12.2)
- Selecting from all possible levels of a statin drug would be an impossible and meaningless experimental design. Explain why.
- For a multiway ANOVA design, when will the differences in the Random versus Fixed Factor make a difference?
Quiz Chapter 14.3
Fixed effects, random effects
Chapter 14 contents
14.2 – Sources of variation
Introduction
Sources of variation, or components of the two-way ANOVA include two factors, each with two or more levels (groups), and collectively, factors are often referred to as the main effects in these types of ANOVA. The other source of variation in a two-way ANOVA is the interaction between the two factors. Below, I have listed the important components, although I have not included how the sum of squares are calculated. You are expected to know the sources of variation for this most basic two-way ANOVA table (Fig. 1). You should also be able to solve any missing elements in one of these tables by utilizing any included information.
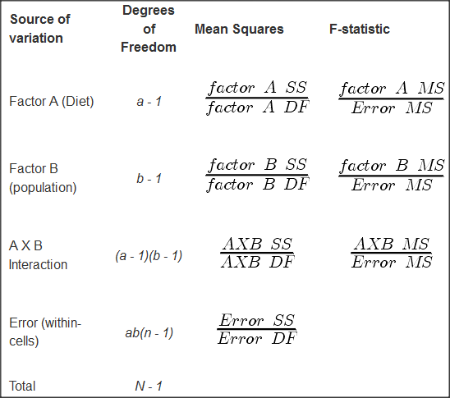

Figure 1. ANOVA table for two-way, balanced, replicated design.
Taking each row from Figure 1 one at a time we have
| Source | DF | Mean Squares | F-statistic |
| First Factor |
where Source refers to the source of variation, DF refers to Degrees of Freedom, a is the number of levels (groups) of the first factor, SS refers to Sum of Squares, and MS refers to the Mean Squares.
Next is the second factor
| Source | DF | Mean Squares | F-statistic |
| First Factor |
where b is the number of levels (groups) of the second factor. Next is the interaction between the first and second factors.
| Source | DF | Mean Squares | F-statistic |
| Interaction |
and lastly the Within-cell Error or residual source of variation
| Source | DF | Mean Squares | F-statistic |
| Error |
where n is the number of experimental units for each group. Note that if the sample size differs for one or more groups (levels),then the design would be unbalanced and this formula does not work to determine the degrees of freedom. The total degrees of freedom for the two-way ANOVA is simply N – 1, where N is the sample size for the entire problem; a little algebra shows that N may be calculated as
![]()
Unbalanced designs
An unbalanced design implies that observations are missing value for one or more groups. What to do if data are missing? Decision depends on how the data are missing (see Chapter 5). For example, if data are missing at random with respect to treatment, then this should not affect inference. If data are missing not at random, then inference, logically, must be impacted. Calculating the ANOVA, moreover, becomes a different matter. In the one-way ANOVA, no real problem arises although setting up contrasts among the levels requires a weighting term to be factored into the calculations. For higher-level ANOVA involving two or more factors the sums of squares for treatment effects are no longer simple partitioning into the different sources of variation. The sources overlap and the order by which the Factors enter into the statistical model now affects the calculations. Thus, while setting up the calculations for the balanced design is straight-forward, perhaps surprisingly, if group sizes differ, this simple relationship for calculating the degrees of freedom, sums of squares, and Mean squares become an unsolvable problem. This problem is largely solved by the general linear model.
Questions
- Based on the results of a two-way ANOVA, the error sums of squares (SSE) was computed to be 160. If we ignore one of the factors and perform a one-way ANOVA using the same data, will the SSE be the same as in the two-way ANOVA, or will it increase? Decrease? Explain your choice.
- While conducting a two-way ANOVA, you conclude that a statistically significant interaction exists between factor 1 and factor 2. What should be your next step? Do you drop the interaction term from the model and redo the analysis or do you report the results of factor effects including the non-significant interaction?
Quiz Chapter 14.2
Sources of variation
Chapter 14 contents
14.1 – Crossed, balanced, fully replicated designs
Introduction
“Biology is complicated” (p. 25, National Research Council [2005]), and as researchers we need to balance our need for statistical models that fit the data well and provide insight into the phenomenon in question against compressing that complexity into ways that do not reflect the phenomenon or hinder further progress in understanding the phenomenon. From our view as researchers then, we recognize that an experiment with only one causal variable is not likely to be informative. For example, while diet has a profound effect on weight, clearly, activity levels are also important. At a minimum, when considering a weight loss program, we would want to control or monitor activity of the subjects. This is a two-factor model, the two factors diet and activity, are expected to both affect weight loss, and, perhaps, they may do so in complicated ways (e.g., on DASH diet, weight loss is accelerated when subjects exercise regularly).
Before we proceed, a word of caution is warranted. Prior to the 1990s, one could be excused for implementing experiments with simple designs that are suitable for analysis by contingency tables, t-tests, and one-way ANOVA. Now, with powerful computers available to most of us, and the feature-rich statistical packages installed on these computers, we can do much more complicated analyses on, hopefully, more realistic statistical models. This is surely progress, but caution is warranted nonetheless — just because you have powerful statistical tests available does not mean that you are free to use them — there is much to learn about the error structures of these more complicated models, for example, and how inferences are made across a model with multiple levels of interaction. In general it is preferred that experimental researchers consult and work with knowledgeable statisticians so that the most efficient and powerful experiment can be designed and subsequently analyzed with the correct statistical approach (Quinn and Keough 2002). Our introductory biostatistics textbook is not enough to provide you with all of the tools you would need and while I do advocate self-learning when it comes to statistics I do so provided we all agree that we are likely not getting the full picture this way. What we can do is provide an introduction to the field of experimental design with examples of classical designs so that the language and process of experimental design from a statistical point of view will become familiar and allow you to participate in the discussion with a statistician and read the literature as an informed consumer.
Two-factor ANOVA with replication
Our one factor statistical models can easily be extended to reflect more complicated models of causation, from one factor to two or more. We begin with two factors and the two-way ANOVA. Now we want to extend our discussion to examine how we can analyze data where we have two factors that may cause variation in the one response variable.
Consider the following two way data set.
| Diet A Population 1 |
Diet A Population 2 |
Diet B Population 1 |
Diet B Population 2 |
| 4 | 5 | 12 | 5 |
| 6 | 8 | 15 | 7 |
| 5 | 9 | 11 | 8 |
I’ve included the stacked version of this dataset at the end of this page (scroll to end or click here).
Question: What is the response variable? Which variable is the Factor variable? What are the classes of treatments and the levels of the treatments?
Answer.
Factors: Diet & Population
Levels: A, B for Diet;
Observations from population 1 or 2
Note the replication: for every level of Diet (A or B) there is an equal number of individuals from the 2 populations. Said another way, there are three replicates from population 1 for Diet A, 3 replicated from population 2 for Diet A, etc.
And finally, we say that the experiment is CROSSED: Both levels of Diet have representatives of both levels of Population.
In order to properly analyze this type of research design (2 factor ANOVA, with equal replication), the data must be crossed. “Crossed” means that each level of Factor 1 must occur in each level of Factor 2.
From the example above: each population must have individuals given diet A and diet B.
Each of the collection of observations from the same combination of Factor 1 and Factor 2 is called a CELL:
All individuals in Diet A and Population 1 are in cell 1.
All individuals in Diet A and Population 2 are in cell 2.
All individuals in Diet B and Population 1 are in cell 3.
All individuals in Diet B and Population 2 are in cell 4.
If the data is completely crossed then you can calculate the number of cells:
Number of Levels in Factor 1 x Number of Levels in Factor 2 = Total Number of Cells
From the above example: 2 Diets x 2 Populations = 4 cells.
How to analyze two factors?
One solution (but inappropriate) is to do several separate One-Way ANOVAs.
There are two reasons that this approach is not ideal:
- This approach will increase the number of tests performed and therefore will increase the chance of rejecting a Null Hypothesis when it is true (increase our p value without us being aware that it is changing – R and Rcmdr will not tell you there is a problem). This is analogous to the problems that we have seen if we perform multiple t-tests instead of a Multi-Sample ANOVA.
- More importantly, there may be interactions among the TWO Factors in how they effect the response variable. One of the more interesting possible outcomes is that the influence of one of the Factors DEPENDS on the second FACTOR. In other words, there is an interaction between factor one and factor two on how the organism responds.
Here is a graph that illustrates one possible outcome:
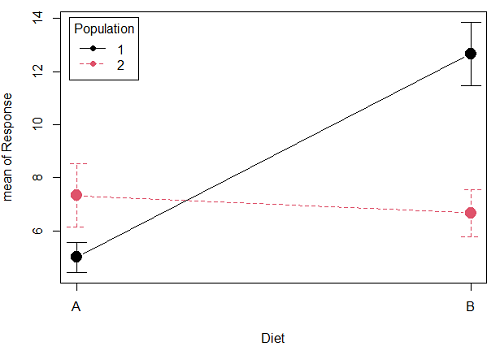
Figure 1A. One of several possible outcome of two treatments (factors). A clear interaction: First Diet level population 1 has greatest weight change, whereas for second diet level, population 2 has greatest weight change.
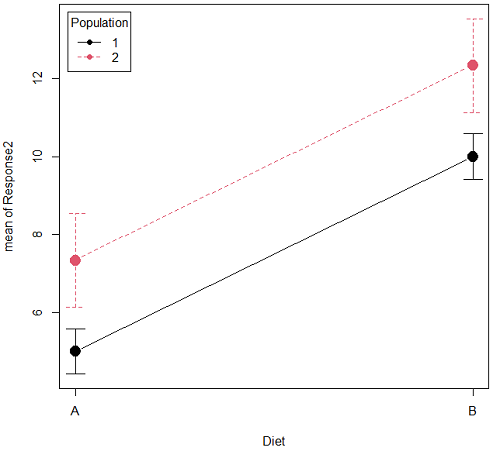
Figure 1B. One of several possible outcome of two treatments (factors). Clearly, no interaction: Population 1 always lower response than Population 2 regardless of Diet.
R code for plots
Rcmdr: Graphs → Plot of means… then added pch=19 and modified legend.pos= from "farright" to "topleft".
Figure 1A.
with(pops2, plotMeans(Response, Diet, Population, pch=19, error.bars="se", connect=TRUE, legend.pos="topleft"))
Figure 1B.
with(pops2, plotMeans(Response2, Diet, Population, pch=19, error.bars="se", connect=TRUE, legend.pos="topleft"))
Figure 1A and 1B shows that BOTH factors, Diet and Population, effect the Response of the subjects. Figure 1A also shows that the effects across Diet are not consistent: the responses are different. Individuals in Population 1 show decreased change in weight going from Diet A(1) to Diet B (2). But, individuals from Population 2 do just the opposite.
Figure 1A, because the effect of Diet cannot be interpreted without knowing which population you’re looking at, this is called an interaction between Factor 1 and Factor 2. It’s the part of the variation in the response NOT accounted for by either factor.
We can see the importance of doing the two-factor ANOVA by showing what would happen if we did two One-Factor (one-way) ANOVAs. For the first One-Factor (multi-sample) ANOVA we can examine the effect of Diet on weight. We could do this by combining the individuals from populations 1 & 2 that are given diet A (Diet A group) and then combining individuals from populations 1 & 2 that are given diet B (Diet B group).
An incorrect analysis of a two-way designed experiment
Statistical software will do exactly what you tell it to do, therefore, there is nothing to stop you from analyzing your two factor experimental design one variable at a time. It is statistical wrong to do so, but, again, there is nothing in the software that will prohibit this. So, we need to show you what happens when you ignore the experimental design in favor of a simple application of statistical analysis.
First, take a look at our two-way example with Diet as a factor and Population as another factor.
Here’s is the one-way ANOVA for Diet only.
aov(Response ~ Diet, data=pops)
| One-way ANOVA table (ignoring the other factor) |
|||||
| Source | DF | Sum of Squares | Mean Squares | F | P |
| Diet | 1 | 36.75 | 36.75 | 4.26 | 0.066 |
| Error | 10 | 86.17 | 8.62 | ||
| Total | 11 | 122.92 | |||
When we ignore (combine) the identity of the two populations in this example we see that it would APPEAR that Diet has NO EFFECT on the weight of the individuals, at least based on our statistical significance cut-off of Type I error set to 5%. Similarly, if we ignore Diet and compare responses by Population, p-value was 0.367, not statistically significant (confirm p-value from one-way ANOVA on your own).
Now let’s do the analysis correctly and pay attention to the main effect Diet.
Here’s the 2-way ANOVA table.
lm(Response ~ Diet*Population, data=pops, contrasts=list(Diet ="contr.Sum", Population + ="contr.Sum"))
| Two-way ANOVA (the correct analysis!) | |||||
| Source | DF | SS | MS | F | P |
| Diet | 1 | 36.75 | 36.75 | 12.25 | 0.008 |
| Population | 1 | 10.08 | 10.08 | 3.36 | 0.104 |
| Interaction | 1 | 52.08 | 52.08 | 17.36 | 0.003 |
| Error | 8 | 24.00 | 3.00 | ||
| Total | 11 | 122.92 | |||
We can visualize the results by plotting the means for each treatment group (Fig. 2).
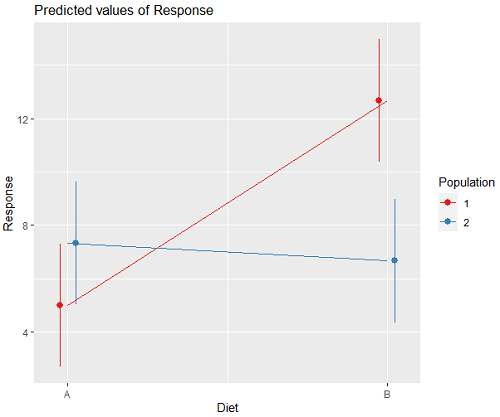
Figure 2. Plots of the main effects for Diet factor, levels A and B, and Population, levels 1 and 2.
R code for plot Fig 2A.
library(sjPlot)
library(sjmisc)
library(ggplot2)
plot_model(LinearModel.1, type = "pred", terms = c("Diet", "Population")) + geom_line()
And then for the interaction (Fig. 3).
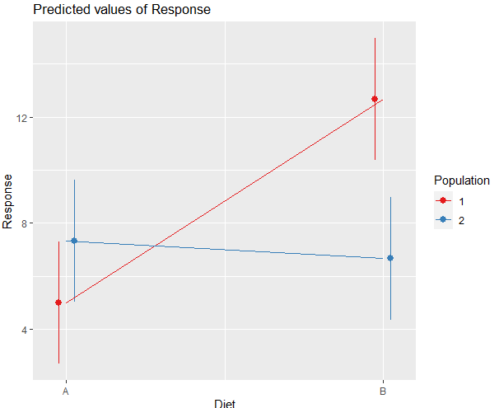
Figure 3. Interaction plot between two factors, Diet and Population.
R code: two-way ANOVA
The more general approach to running ANOVA in R is to use the general linear model function, lm(), saved as object MyLinearModel.1, for example, then follow up with
Anova(MyLinearModel.1, type="II")
to obtain the familiar ANOVA table. The lm() menu is obtained in Rcmdr by following Statistics→ Fit models→ Linear model…, and entering the model (Fig. 4). In this case, the model was
![]()
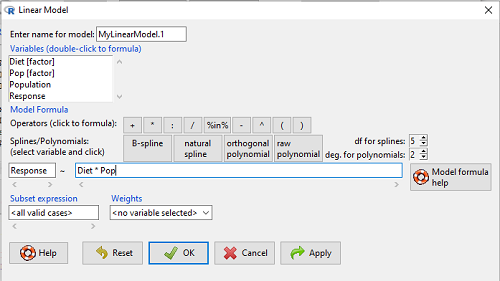
Figure 4. Linear model menu in Rcmdr.
Output from lm() function for this example LinearModel.2 <- lm(Response ~ Diet * Pop, data=pops) summary(LinearModel.2) Call: lm(formula = Response ~ Diet * Pop, data = pops) Residuals: Min 1Q Median 3Q Max -2.3333 -1.1667 0.1667 1.0833 2.3333 Coefficients: Estimate Std. Error t value Pr(>|t|) (Intercept) 5.000 1.000 5.000 0.00105 ** Diet[T.B] 7.667 1.414 5.421 0.00063 *** Pop[T.2] 2.333 1.414 1.650 0.13757 Diet[T.B]:Pop[T.2] -8.333 2.000 -4.167 0.00314 ** --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 Residual standard error: 1.732 on 8 degrees of freedom Multiple R-squared: 0.8047, Adjusted R-squared: 0.7315 F-statistic: 10.99 on 3 and 8 DF, p-value: 0.003285
We want the ANOVA table, so run
Anova(MyLinearModel.1, type="II")
or in Rcmdr, Models → Hypothesis tests → ANOVA table… Accept the defaults (Types of tests = Type II, uncheck use of sandwich estimator), and press OK. I’ll leave that for you to do (see Questions).
Interaction, explained
How can we visualize the effects of the Factors and the effects of the interaction? Plot the means of a two-factor ANOVA (Fig. 5). An interaction is present if the lines cross (even if they cross outside the range of the data), but if the lines are parallel, no interaction is present.
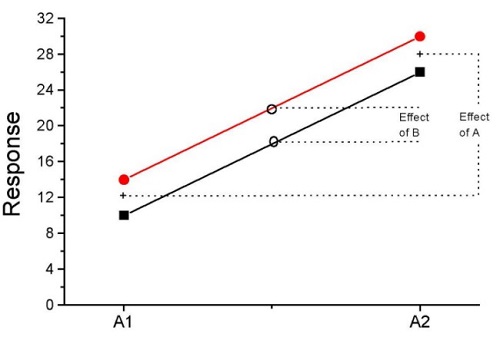
Figure 5. A plot showing no interaction between factor A and factor B for some ratio scale response variable.
A large effect of factor A – compare means
A small effect of factor B – compare means
Little or no interaction – lines are parallel
Three hypotheses for the Two-Factor ANOVA
The important advance in our statistical sophistication (from one to two factors!!) allows us to ask three questions instead of just two question:
- Is there an effect of Factor 1?
- HO: There is no effect of Factor 1 on the response variable.
- HA: There is an effect of Factor 1 on the response variable.
- Is there an effect of Factor 2?
- HO: There is no effect of Factor 2 on the response variable.
- HA: There is an effect of Factor 2 on the response variable.
- Is there an INTERACTION between Factor 1 & Factor 2?
- HO: There is no interaction between Factor 1 & Factor 2 on the response variable.
- HA: There is an interaction between Factor 1 & Factor 2 on the response variable.
Questions
- In the crossed, balanced two-way ANOVA, how many Treatment groups are there if Factor 1 has three levels and Factor 2 has four levels?
A. 3
B. 4
C. 7
D. 9
E. 12 - What is meant by the term “balanced” in a two-way ANOVA design?
A. Within levels of a factor, each level has the same sample size
B. Each level of one factor occurs in each level of the other factor
C. There are no missing levels of a factor.
D. Each level of a factor must have more than one sampling unit. - What is meant by the term “crossed” in a two-way ANOVA design?
A. Within levels of a factor, each level has the same sample size
B. Each level of one factor occurs in each level of the other factor
C. There are no missing levels of a factor.
D. Each level of a factor must have more than one sampling unit. - What is meant by the term “replicated” in a two-way ANOVA design?
A. Within levels of a factor, each level has the same sample size
B. Each level of one factor occurs in each level of the other factor
C. There are no missing levels of a factor.
D. Each level of a factor must have more than one sampling unit. - Use the multi-way ANOVA command in
Rcmdrto generate the ANOVA table for the example data set. - Use the linear model function and Hypothesis tests in
Rcmdrto generate the ANOVA table for the example data set.
Quiz Chapter 14.1
Crossed, balanced, fully replicated designs
Data set
Don’t forget to convert the numeric Population variable to character factor, e.g., a new object called Pop. The R command is simply
Pop <- as.factor(Population)
But easy to use Rcmdr also. From within Rcmdr select Data → Manage variables in active dataset → Convert numeric variables to factors…
| Diet | Population | Response |
| A | 1 | 4 |
| A | 1 | 6 |
| A | 1 | 5 |
| A | 2 | 5 |
| A | 2 | 8 |
| A | 2 | 9 |
| B | 1 | 12 |
| B | 1 | 15 |
| B | 1 | 11 |
| B | 2 | 5 |
| B | 2 | 7 |
| B | 2 | 8 |
Chapter 14 contents
14 – ANOVA designs, multiple factors
Introduction
In our previous discussions about t-tests and ANOVA we focused on procedures with one dependent (response) variable and a single independent (predictor) factor variable that may cause variation in the response variable. In this chapter we extend our discussions about the general linear model by reviewing
- One-way ANOVA and provided a few examples of the one-way design.
- To review and set the stage for adding a second independent variable to the model.
Additional one-way ANOVA examples
1. In a plants, we may have a response variable like height and one factor variable (location: sun vs. shade) thought to influence plant height (eg, Aphalo et al 1999).
2. Pulmonary macrophage phagocytosis behavior (response variable) after exposure of toads to clean air or ozone (factor with 2 levels) (Dohm et al. 2005).
3. Monitor weight change on subjects after 6 weeks eating different diet (DASH, control) (Elmer et al. 2006).
All three of the examples are based on the same statistical model which may be written as:
![]()
where μ is the grand mean, Y is the response variable and A is the independent variable, or factor, with k = 1, 2, …. K levels, groups, or treatments. The total number of experimental units (eg, subjects) is given by i = 1, 2, 3, … n. Note that in the first and third examples, because there were only two groups (example 1: k = location, shade; example 3: k = DASH, control) note that this problem could have been evaluated as an independent sample t-test. For the second example, there were three groups so k = clean air, first ozone level, second ozone level).
Two-way ANOVA with replication
Biology experiments are typically more complicated than a single t-test or one-way ANOVA design can handle; rarely would we conduct an experiment that reflects only one source of variation.
For example, while diet has a profound effect on weight, clearly, activity levels are also important. At a minimum, when considering a weight loss program, we would want to control or monitor activity of the subjects. This is a two-factor model, and the main effects, the two factors, were diet (factor A) and activity, (factor B). Both are expected to affect weight loss, and, perhaps, they may do so in complicated ways — an interaction (eg, on DASH diet, weight loss is accelerated when subjects exercise regularly).
![]()
The subject of this chapter is the introduction to two-way ANOVA designs. In fact, to many, ANOVA design is practically synonymous to a statistician when they think about experimental design (Lindman 1992; Quinn and Keough 2002). As noted by Quinn and Keough (2002) in the preface to their book, “… many biological hypotheses, even deceptively simple ones, are matched by complex statistical models” (p. xv). Once you start adding factor variables there becomes a number of ways in which the groups and experimental units can be distributed, and thus impact the inferences one can make from the ANOVA results. The first statistical model we introduced was the one-way ANOVA. Next, we begin the two-way ANOVA with the crossed, balanced, fully replicated design. Along the way we introduce model symbols to help us communicate the design structure and implications of the statistical models.
Quizzes in this chapter
A total of 71 questions among the several subchapters, a mix of true or false and multiple choice question format.
Chapter 14 contents
13.4 – Tests for Equal Variances
Introduction
In order to carry out statistical tests correctly, we must test our data first to see if our sample conforms to the assumptions of the statistical test. If our data do not meet these assumptions, then inferences drawn may be incorrect. How far off our inferences may be depends on a number of factors, but mostly it depends on how far from the expectations our data are.
One assumption we make with parametric tests involving ratio scale data is that the data could be from a normally distributed population. The other key assumption introduced, but not described in detail for the two-sample t-test, was that the variability in the two groups must be the same, i.e., homoscedasticity. Thus, in order to carry out the independent sample t-test, we must assume that the variances are equal.
There are two general reasons we may want to concern ourselves with a test for the differences between two variances
- The t-test (and other tests like one-way ANOVA) requires that the two samples compared have the same variances. If the Variances are Not Equal we need to perform a modified t-test (see Welch’s formula).
- We may also be interested in the differences between the variances in two populations.
Example 1: In genetics we might be interested in the difference between the variability of response of inbred lines (little genetic variation but environmental variation) versus an outbred populations (lots of genetic and environmental variation).
Example 2: Environmental stress can cause organisms to have developmental instability. This might cause organisms to be more variable in morphology or the two sides (right & left) of an organism may develop non-symmetrically. Therefore, polluted environments might cause organisms to have greater variability compared to non-polluted environments.
The first way to test the variances is to use the F test. This works for two groups.
![]()
For more than two groups, we’ll use different tests (e.g., Bartlett’s test, Levene’s test).
Remember that the formula for the sample variance is
![]()
The Null Hypothesis is that the two samples have the same variances:
![]()
The Alternative Hypothesis is that the two samples do not have the same variances:
![]()
Note: I prefer to evaluate this as a one-tailed test: identify the larger of the two variances and take that as the numerator Then, the null hypothesis is that
![]()
and therefore, the alternative hypothesis is that (i.e., a one-tailed test).
![]()
Another way to state equal variance test is that we are testing for homogeneity of variances. You may run across the term homoscedasticity; it is the same thing, just a different three dollar word for “equal variances.”
Stated yet another way, if we reject the null hypothesis, then the variances are unequal or show heterogeneity. An additional and equivalent $3 word for inequality of variances is called heteroscedasticity.
More about the F-test
For the F-test, the null hypothesis is that the variances are equal. This means that the “expected” F value will be one: F = 1.0. (The F-distribution differs from t-distribution because it requires 2 values for DF, and ranges from 1 to infinity for every possible combination of v1 and v2).
To evaluate the null hypothesis we need the degrees of freedom. For the F test we need two different degrees of freedom, one set for each group): from Table 2, Appendix – F distribution, look up 5% Type I error line in this table because we make it one tailed.
I need the F-test statistic at ![]()
Examples of difference between two variances, Table 1.
Sample 1: Aggressiveness of Inbred Mice (number of bites in 30 minutes)
Sample 2: Aggressiveness of Outbred Mice (number of bites in 30 minutes)
Table 1. Aggression by inbred and outbred mice.
| Sample 1
Aggressiveness of Inbred Mice |
Sample 2
Aggressiveness of Outbred Mice |
| 3 | 4 |
| 5 | 10 |
| 4 | 4 |
| 3 | 7 |
| 4 | 7 |
| 5 | 10 |
| 4 = mean | 7 = mean |
- Identify the null and alternative hypotheses
- Calculate Variances
- Calculate F-test
- The “test statistic” for this hypothesis test was F = 7.2 / 0.8 = 9.0
- Determine Critical Value of the F table (Table 2, Appendix – F distribution)
Example of how to find the critical values of the F distribution for ![]() and numerator
and numerator ![]() and denominator
and denominator ![]() .
.
Table 2. Portion of F distribution, see Appendix – F distribution.
| α = 0.05 | v1 | ||||||||
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | ||
| v2 | 1 | 230 | |||||||
| 2 | 19.3 | ||||||||
| 3 | 9.01 | ||||||||
| 4 | 6.26 | ||||||||
| 5 | 6.61 | 5.79 | 5.41 | 5.19 | 5.05 | 4.95 | 4.88 | 4.82 | |
| 6 | |||||||||
Or, instead of using tables, use R
Rcmdr: Distributions → F distribution → F probabilities
and enter the numbers as shown below (Fig 1).
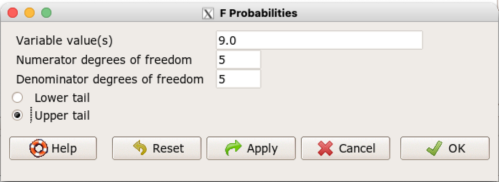
Figure 1. Screenshot R Commander F distribution probabilities.
This will return the p-value, and you would interpret this against your Type I error rate of 5% as you do for other tests.
From Table 2, Appendix – F distribution we find ![]()
And the p-value = 0.015
pf(c(9), df1=5, df2=5, lower.tail=FALSE) [1] 0.01537472
Question: Reject or Accept null hypothesis?
Question: What is the biological interpretation or conclusion from this result?
R code
Rather than play around with the tables of critical values, which are awkward and old-school (and I am showing you these stats tables so that you get a feel for the process, not so you’d actually use them in real practice), use Rcmdr to generate the F test and therefore return the F distribution probability value. As you may expect, R provides a number of options for testing the equal variances assumption, including the F test. The F test is limited to only two groups and, because it is a parametric test, it also makes the assumption of normality, so the F test should not be viewed as necessarily the best test for the equal variances assumption among groups. We present it here because it is a logical test to understand and because of its relevance to the Mean Square ratios in the ANOVA procedures.
So, without further justification, here is the presentation on how to get the F test in Rcmdr. At the end of this section I present a better procedure than the F test for evaluating the equal variance assumption called the Levene test.
Return to the bite data in the table above and enter the data into an R data frame. Note that the data in the table above are unstacked; R expects the data to be stacked, so either create a stacked worksheet and transcribe the data appropriately into the cells of the worksheet, or, go ahead and enter the values into two separate columns then use the Stack variables in active data set… command from the Data menu in Rcmdr.
Then, proceed to perform the F test.
Rcmdr: Statistics → Variances → Two variances F-test…
The first context menu popup is where you enter the variables (Fig 2).
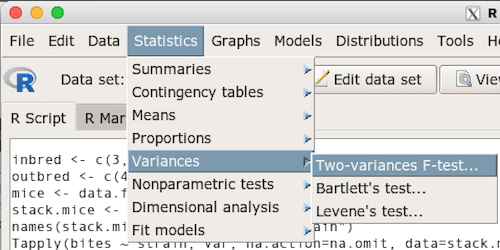
Figure 2. Screenshot data options R Commander F test
Because there are only two variables in the data set and because Strain contains the text labels of inbred or outbred whereas the other variable is numeric data type, R will correctly select the variables for you by default. Select the “Options” tab to set the parameters of the F test (Fig. 3).
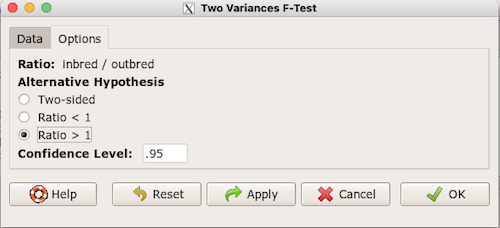
Figure 3. Screenshot menu options R Commander F test.
When you are finished setting the alternative hypothesis and confidence levels, proceed with the F test by clicking the OK button.
F test to compare two variances data: mice.aggression F = 0.1111, num df = 5, denom df = 5, p-value = 0.03075 alternative hypothesis: true ratio of variances is not equal to 1 95 percent confidence interval: 0.01554788 0.79404243 sample estimates: ratio of variances 0.1111111
End of R output.
Levene’s test of equal variances
We will discuss this test in more detail following our presentation on ANOVA. For now, we note that the test works on two or more groups and is a conservative test of the equal variance assumption. Nonparametric tests in general make fewer assumptions about the data and in particular make no assumption of normality like the F test. It is in this context that the Levene’s test would be preferable over the F test. Below we present only how to calculate the statistic in R and Rcmdr and provide the output for the same mouse data set.
Assuming the data are stacked, obtain the Levene’s test in Rcmdr by clicking on (Fig 4).
Rcmdr: Statistics → Variances → Levene’s test…
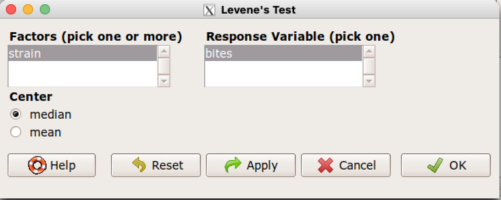
Figure 4. Screenshot menu options R Commander Levene’s test.
Select the median and the factor variable (in our case “Strain”) and the numeric outcome variable (“Bites”), then click OK button.
Tapply(bites ~ strain, var, na.action=na.omit, data=mice.aggression) # variances by group inbred outbred 0.8 7.2 leveneTest(bites ~ strain, data=mice.aggression, center="median") Levene's Test for Homogeneity of Variance (center = median) Df F value Pr(>F) group 1 4 0.07339 . 10 --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
End of R output
Compare the p-values. In both cases — F test and Levene’s test — we’re testing the same hypothesis (equal variances), but the p-values do not agree between the parametric F test and the nonparametric Levene’s test! If we were to go by the results of the F test, p-value was 0.031, less than out Type I error of 5% and we would tentatively conclude that the assumption of equal variances may not apply. On the other hand, if we go with the Levene’s test the p-value was 0.074, which is greater than our Type I error rate of 5% and we would therefore conclude the opposite, that the assumption of equal variances might apply! Both conclusions can’t hold, so which test result of equal variances do we prefer, the parametric F test or the nonparametric Levene’s test? Answer — we’d go with Levene’s because the F test is parametric and, therefore, also assumes normality.
Cahoy’s bootstrap method
Draft. Cahoy (2010). Variance-based statistic bootstrap test of heterogeneity of variances. we discuss bootstrap methods in Chapter 19.2; in brief, bootstrapping involves resampling the dataset and computing a statistic on each sample. This method may be more powerful, that is, more likely to correctly reject the null hypothesis when warranted, compared to Levene’s test.
for now, install package testequavar.
Function for testing two samples.
equa2vartest(inbred, outbred, 0.05, 999)
R output
[[1]] [1] "Decision: Reject the Null" $Alpha [1] 0.05 $NumberOfBootSamples [1] 999 $BootPvalue [1] 0.006
The output “Decision: Reject the Null” reflects output from a box-type acceptance region.
Compare results from Levene’s test: p-value 0.07339 suggests accept hypothesis of equal variances, whereas bootstrap method indicates variances heterogenous, i.e., reject equal variance hypothesis. However, re-running the test without setting the seed for R’s pseudorandom number generator will result in different p-values. For example, I re-ran Cahoy’s test five times with the following results:
0.008 0.006 0.002 0.01 0.004
Questions
- Test assumption of equal variances by Bartlett’s method and by Levene’s test on
OliveMomentvariable from the Comet tea data set introduced in Chapter 12.1. Do the methods agree? If not, which test result would you choose?- BONUS. Retest homogeneous variance hypothesis by
equa3vartest(Copper.Hazel, Copper, Hazel, 0.05, 999). Reject or fail to reject the null hypothesis by bootstrap method?
- BONUS. Retest homogeneous variance hypothesis by
- Test assumption of equal variances by Bartlett’s method and by Levene’s test on
Heightfrom the O’hia data set introduced in Chapter 12.7. Do the methods agree? If not, which result would you choose?- BONUS. Retest homogeneous variance hypothesis by
equa3vartest(M.1, M.2, M.3, 0.05, 999). Reject or fail to reject the null hypothesis by bootstrap method?
- BONUS. Retest homogeneous variance hypothesis by
Quiz Chapter 13.4
Tests for Equal Variances
R code reminders
The bootstrap method expects the variables in unstacked format. A simple method to extract the variables from the stacked data is to use a command like the following. For example, extract OliveMoment values for Copper-Hazel treatment.
Copper.Hazel <- cometTea$OliveMoment[1:10]
For Copper, Hazel, replace above with [11:20], and [21-30] respectively. Note: changed variable name from Copper-Hazel to Copper.Hazel. The hyphen is a reserved character in R.
Chapter 13 contents
13.3 – Test assumption of normality
Introduction
I’ve commented numerous times that your conclusions from statistical inference are only as good as the validity of making the and applying the correct procedures. This implies that we know the assumptions that go into the various statistical tests, and where possible, we critically test the assumptions. From time to time then I will provide you with “tests of assumptions.”
Here’s one. The assumption of normality, that your data were sampled from a population with a normal distribution for the variable of interest, is key and there are a number of ways to test this assumption. Hopefully as you read through the next section you can extend the logic to any distribution; if the data are presumed to come from a binomial, or a Poisson, or a uniform distribution, then the logic of goodness of fit tests would apply.
How to test normality assumption
It’s not a statistical test per se, but the best is to simply plot (histogram) the data. You can do these graphics by hand, or, install the data mining package rattle which will generate nice plots useful for diagnosing normality issues.
Note 1: rattle (R Analytic Tool To Learn Easily) is a great “data-mining” package. We introduced the package in Chapter 3 — Exploring data.
The rattle histogram plot (Fig 1) superimposes a normal curve over the data, which allows you to “eyeball” the data.
First, the eye test. I used the R-package rattle for this on a data set of comet tail lengths of rat lung cells exposed to different amounts of copper in growth media (scroll to bottom of page or click here to get the data).
In addition to the histogram (Fig 1 top image), I plotted the cumulative function (Fig 1 bottom image). In short, if the data set were normal, then the cumulative frequency plot should look like a straight line.
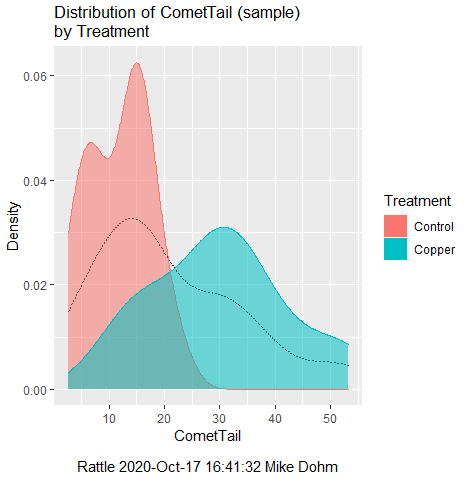
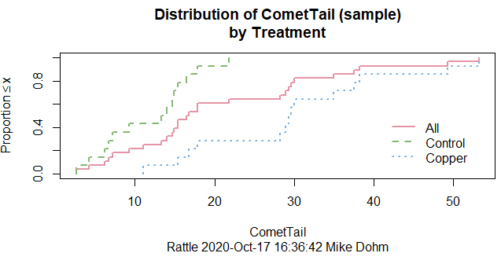
Figure 1. Rattle descriptive graphics on Comet Copper dataset. Dotted line (top image) and red line (bottom image) follow the combined observations regardless of treatment.
So, just looking at the data set, we don’t see clear evidence for a normal-like data set. The top image (Fig 1) looks stacked to the left and the cumulative plot (bottom image) is bumped in the middle, not falling on a straight line. We’ll need to investigate the assumption of normality more for this data set. We’ll begin by discussing some hypothetical points first, then return to the data set.
Goodness of fit tests of normality hypothesis
While graphics and the “eyeball test” are very helpful, you should understand that whether or not your data fits a normal distribution, that’s a testable hypothesis. The null hypothesis is that your sample distribution fits are normal distribution. In general terms, these “fit” hypotheses are viewed as “goodness of fit” tests. Often times, the test is some variation of a ![]() problem: you have your observed (sample distribution) and you compare it to some theoretical expected value (e.g., in this situation, the normal distribution). If your data fit a normal curve, then the test statistics will be close to zero.
problem: you have your observed (sample distribution) and you compare it to some theoretical expected value (e.g., in this situation, the normal distribution). If your data fit a normal curve, then the test statistics will be close to zero.
We have discussed before that the data should be from a normal distributed population. To the extent this is true, then we can trust our statistical tests are performing the way they are expected to do. We can test the assumption of normality by comparing our sample distribution against a theoretical distribution, the normal curve. I’ve shown several graphs in the past that “looked normal”. What are the alternatives for unimodal (single peak) distributions (Fig 2)?
Kurtosis describes the shape of the distribution, whether it is stacked up in the middle (leptokurtosis), or more spread out and flattened (platykurtosis).
Skewness describes differences from symmetry about the middle. Left skew the tail of the distribution extends to the left, ie, smaller values.
“Leptokurtosis”
“Platykurtosis”
Negative skew, left skewed
Positive skew, right skewed
Figure 2. Graphs describing different distributions. From top to bottom: Leptokurtosis, platykurtosis, negative skew, positive skew.
The easiest procedures for goodness of fit tests of normality are based on the ![]() distribution and yield a “goodness of fit” test for normal distribution. We discussed the
distribution and yield a “goodness of fit” test for normal distribution. We discussed the ![]() distribution in Chapter 6.9, and used the test in Chapter 9.1.
distribution in Chapter 6.9, and used the test in Chapter 9.1.
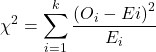
where O refers to the observed data (what we’ve got) and E refers to the expected (e.g., data from a normal curve with same mean and standard deviation as our sample).
To illustrate, I simulated a data set in R.
Rcmdr: Distributions → Continuous distributions → Normal distribution → Sample from normal distribution
I created 100 values, stored them in column 1, and set the population mean = 125 and population standard deviation = 10. And therefore the population standard error of the mean was 1.0.
The resulting MIN/MAX was from 99.558 to 146.16; the sample mean was 124.59 with a sample standard deviation of 9.9164. And therefore the sample standard error of the mean was 0.9916.
Question: After completing the steps above, your data will be slightly different from mine… Why?
But getting back to the main concept, Does our data agree with a normal curve? We have discussed how to construct histograms and frequency distributions.
Let’s try six categories (why six? we discussed this when we talked about histograms).
All chi-square tests are based on categorical data, so we use the counts per category to get our data. Group the data, then count the number of OBSERVED in each group. To get the EXPECTED values, use the Z-score (normal deviate) with population mean and standard deviation as entered above.
Table 1. Tabulated values for test of normality.
| Number of observations | Weight | Normal deviate (Z) | Expected Proportion | Expected number | (Obs – Exp)2 / Exp |
| 105 | 3 | less than or equal to -2 | 0.0228 | 2.28 | 0.227368421 |
| 105 < 115 | 17 | between -1 & -2 = 0.1587 – 0.0228 | 0.1359 | 13.59 | 0.855636497 |
| 115 < 125 | 34 | between -0 & -1 = 0.5 – 0.1587 | 0.3413 | 34.13 | 0.000495166 |
| 125 < 135 | 30 | between +0 & +1 = 0.5 – 0.1587 | 0.3413 | 34.13 | 0.499762672 |
| 135 < 145 | 15 | between +1 & +2 = 0.9772 – 0.8413 | 0.1359 | 13.59 | 0.146291391 |
| > 145 | 1 | greater than or equal to +2 = 1 – 0.9772 | 0.0228 | 2.28 | 0.718596491 |
Then obtain the critical value of the ![]() with df = 6 – 1 = 5 (see Appendix, Table of chi-square critical values , critical
with df = 6 – 1 = 5 (see Appendix, Table of chi-square critical values , critical ![]() = 11.1 with df = 5).
= 11.1 with df = 5).
Thus, we would not reject the null hypothesis and would proceed with the assumption that our data could have come from a normally distributed population.
This would be an OK test, but different approaches, although based on a chi-square-like goodness of fit, have been introduced and are generally preferred. We have just shown how one could use the chi-square goodness of fit approach to testing whether your data fit a normal distribution. A number of modified tests based on this procedure have been proposed; we have already introduced and used one (Wilks-Shapiro), which is easily accessed in R.
Rcmdr: Summaries → Wilks-Shapiro
Like Wilks-Shapiro, another common “goodness-of-fit” test of normality is the Anderson-Darling test. This test is now included with Rcmdr, but it’s also available in the package nortest. After the package is installed and you run the library (you should by now be able to do this!), then at the R prompt (>) type:
ad.test(dataset$variable)
replacing “dataset$variable” with the name of your data set and variable name.
In the context of goodness of fit, a perfect fit means the data are exactly distributed as a normal curve, and the test statistics would be zero. Differences away from normality increase the value of the test statistic.
How do these tests perform on data?
The histogram of our simulated normal data of 100 observations with mean = 125 and standard deviation = 10 (Fig 3).
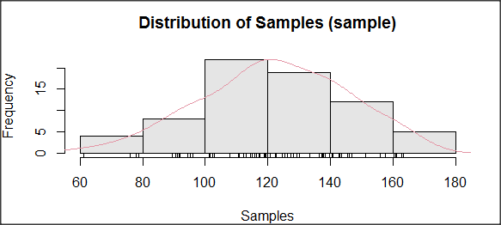
Figure 3. Histogram of simulated normal dataset, μ = 125, σ = 10.
and here’s the cumulative plot (Fig 4).
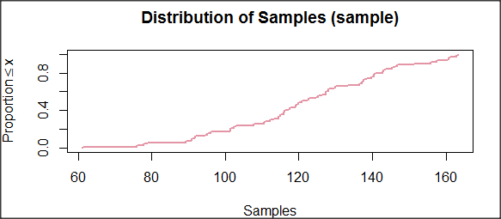
Figure 4. Cumulated frequency of simulated normal dataset, μ = 125, σ = 10.
Results from Anderson-Darling test were A = 0.2491, p-value = 0.7412, where A is the Anderson-Darling test statistic.
Results of the Shapiro-Wilks test on the same data. W = 0.9927, p-value = 0.8716, where W is the Shapiro-Wilks test statistic. We would not reject the null hypothesis in either case because p > 0.05.
Example
Histogram of a data set, highly skewed to the right. 90 observations in three groups of 30 each, with mean = 0 and standard deviation = 1 (Fig 5).
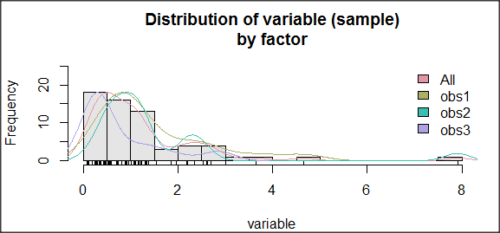
Figure 5. Histogram of simulated normal dataset, μ = 0, σ = 1.
and the cumulative frequency plot (Fig 6).
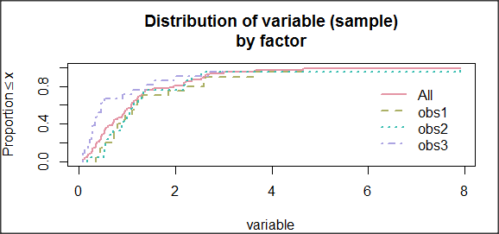
Figure 6. Cumulated frequency of simulated normal dataset, μ = 0, σ = 1.
Results from Anderson-Darling test were A = 9.0662, p-value < 2.2e-16. Results of the Shapiro-Wilks test on the same data. W = 0.6192, p-value = 6.248e-14. Therefore, we would reject the null hypothesis because p < 0.05.
The Shapiro-Wilk test in Rcmdr
Let’s go back to our data set and try tests of normality on the entire data set, i.e., not by treatment groups.
Rcmdr: Statistics → Summaries → Test of normality…
normalityTest(~CometTail, test="shapiro.test", data=CometCopper) Shapiro-Wilk normality test data: CometCopper$CometTail W = 0.91662, p-value = 0.006038
End of R output
Another test, built on the basic idea of a chi-square goodness of fit is the Anderson Darling test. Some statisticians prefer this test and it is one built-in to some commercial statistical packages (e.g., Minitab). To obtain the Anderson-Darling test in R, you need to install a package. After installing nortest package, run the AD test at the command prompt.
require(nortest) ad.test(CometCopper$CometTail) Anderson-Darling normality test data: CometCopper$CometTail A = 1.0833, p-value = 0.006787
End of R output.
Note 2: Because you’ve installed Rcmdr the Anderson-Darling test is available (since version 2.6) without installing the nortest package. You can call Anderson-Darling via the menu system, Statistics → Summaries → Test of normality … or type in the script the following command
normalityTest(~CometTail, test="ad.test", data=CometCopper)
In contrast, Shapiro-Wilks test is part of the base R package, so can be called directly
shapiro.test(CometCopper$CometTail)
P-values are both much less than 0.05, so we would reject the assumption of normality for this data set.
Which test of normality?
Why show you two tests for normality, the Shapiro-Wilks and Anderson-Darling? The simple answer is that both are good as general tests of normality, both are widely used in scientific papers, so just pick one and go with it as your general test of normality.
The more complicated answer is that each is designed to be sensitive to different kinds of departure from normality. By some measures, the Shapiro-Wilks test is somewhat better (i.e., more statistical power to test the null hypothesis), than other tests, but this is not something you want to get into as a beginner. So, I show both of them to you so that you are at least introduced to the concept that there are often more than one way to test a hypothesis. The bottom-line is that plots may be best!
Questions
- Work describe in this chapter involves statistical tests of the assumption of normality. It is just as important, maybe more so, to also apply graphics to take advantage of our built-in pattern recognition functions. What graphic techniques, besides histogram, should be used to view the distribution of the data?
- In R, what command would you use so that you can call the variable name,
CometTail, directly instead of having to refer to the variable asCometCopper$CometTail? - Why are Anderson-Darling, Shapiro-Wilks and other related tests referred to as “goodness of fit” tests? You may wish to review discussion in Chapter 9.1.
- The example tests presented for the Comet Copper data set were conducted on the whole set, not by treatment groups. Re-run tests of normality via
Rcmdr, but this time, select the By groups option and select Treatment.
Quiz Chapter 13.3
Test assumption of normality
Data set used in this page
| Treatment | CometTail |
|---|---|
| Control | 17.86 |
| Control | 16.52 |
| Control | 14.93 |
| Control | 14.03 |
| Control | 13.33 |
| Control | 8.81 |
| Control | 14.70 |
| Control | 9.26 |
| Control | 21.78 |
| Control | 6.18 |
| Control | 9.20 |
| Control | 5.54 |
| Control | 6.72 |
| Control | 2.63 |
| Control | 7.19 |
| Control | 5.39 |
| Control | 11.29 |
| Control | 15.44 |
| Control | 17.86 |
| Contro | l4.25 |
| Copper | 53.21 |
| Copper | 38.93 |
| Copper | 18.93 |
| Copper | 30.00 |
| Copper | 28.93 |
| Copper | 15.36 |
| Copper | 17.86 |
| Copper | 17.50 |
| Copper | 21.07 |
| Copper | 29.29 |
| Copper | 28.21 |
| Copper | 16.79 |
| Copper | 21.07 |
| Copper | 37.50 |
| Copper | 38.22 |
| Copper | 17.86 |
| Copper | 29.64 |
| Copper | 11.07 |
| Copper | 35.00 |
| Copper | 49.29 |
Chapter 13 contents
- Introduction
- ANOVA Assumptions
- Why tests of assumption are important
- Test assumption of normality
- Tests for Equal Variances
- References and suggested readings
13.2 – Why tests of assumption are important
Introduction
Note that R (and pretty much all statistics packages) will calculate t-tests or ANOVA or whatever test you ask for, and return a p-value, but it is up to you to recognize that the p-value is accurate only if the assumptions are met. Thus, you can always estimate a parameter, but interpret its significance (biological, statistical) with caution. The great thing about statistics is that you can directly evaluate whether assumptions hold.
Violation of the assumptions of normality or equal variances can lead to Type I errors occurring more often than the 5% level. That means you will reject the null hypothesis more often than you should! If the goal of conducting statistical tests on results of an experiment is to provide confidence in your conclusions, then failing to verify assumptions of the test are no less important than designing the experiment correctly in the first place. Thus, the simple rule is: know your assumptions, test your assumptions. Evaluating assumptions is a learned skill:
- Conduct proper data exploration.
- Use specialized statistical tests to evaluate assumptions.
- data normally distributed? eg, histogram, Shapiro-Wilk test.
- groups equal variance? eg, box-plot, Levene’s median test.
- Evaluate influence of any outlier observations.
What to do if the assumptions are violated?
This is where judgement and critical thinking apply. You have several options.
First, if the violation is due to only a few observations, then you might proceed anyway, in effect invoking the Central Limit Theorem as justification.
Second, you could check your conclusions with and without the few observations that seem to depart from the trend in the rest of the data set — if your conclusion holds without the “outliers”, then you might conclude that your results are robust).
Third, you might apply a data transform reasoning that the distribution from which the data were sampled was log-normal, for example. Applying a log transform (natural log, base 10, etc,) will tend to make the variances less different among the groups and may also improve normality of the samples.
Fourth, if the violated assumption involves variances differ across different groups or levels of the factors or predictors, then there are more sophisticated alternatives, including use of Generalized Linear Models and creating a model to capture the relationship between predictor means and variances. Additionally, Weighted Least Squares (WLS) can be employed, giving more weight to observations with smaller variances.
Fifth, if a nonparametric test is available, you might use it instead of the parametric test. For example, we introduced the Levene’s test of equal variances as a better choice than the parametric F test. Additionally, there are many nonparametric alternatives to parametric tests. For example, t-tests are parametric and their non-parametric alternatives include tests like Wilcoxon and Mann-Whitney. ANOVA, too is parametric, and its nonparametric alternative version is called rank ANOVA (see also Kruskal-Wallis). See Chapter 15 for nonparametric tests.
Finally, a resampling approach could be taken, where the data themselves are used to generate all possible outcomes like the Fisher Exact test; with large sample size, bootstrap or Jackknife procedures are used (Chapter 19).
For now, let’s introduce you to the kinds of nonparametric statistical tests for which the t-test is just one example. For the independent sample t-test, our first method to account for the possible violation of equal variances is a parametric test, Welch’s variation of the Student’s t-test. Instead of the pooled standard deviation, Welch’s test accounts for each group’s variance in the denominator.
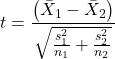
The degrees of freedom for the Welch’s test are now
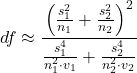
where df for Student’s t-test was ![]() . Note that Welch’s test assumes normal distributed data.
. Note that Welch’s test assumes normal distributed data.
Note 1: In R and therefore R Commander, the default option for conducting the t-test is Welch’s version, not the standard t-test (Fig 1).
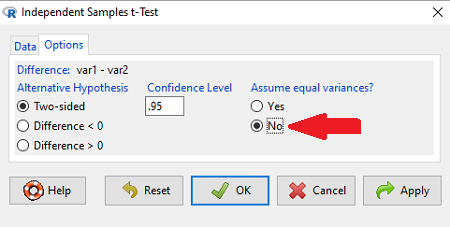
Figure 1. Screenshot of Rcmdr options menu for independent t-test. Red arrow points to default option “No,” which corresponds to Welch’s test.
See discussions in Olsen (2003), Choi (2005), and Hoekstra et al (2012).
How to report choices made about data transformations?
Eventually, with all of the analysis completed, one has to decide what to report. We touched on How to Report Statistics in Chapter 4. However, that chapter emphasized a balance between reporting results in sentences, tables, or as graphics. Here, we need to emphasize a different quality about reporting of statistical methods, that is, transparency. In both reporting contexts — how to report the results and how to report any data transformations — the goal is to provide sufficient detail so that the reader can follow (reproducibility criteria). If guidance presented on Chapter 13.1 and on this page is taken literally, there would seem to be a nearly endless number of graphs and statistical results to report. Clearly, this can’t be the answer.
For students in BI311 there should be clear guidance for what is expected for reporting in homework. More generally, when analysis leads to a completed project ready for reporting at conference or as a journal article, then the goal is to provide enough information so that the reader can evaluate the choices you made.
Consider the body mass data presented on this page. Results of various tests on both raw and log10-transformed data were provided. The context was check of normality assumption, and, based on the results we should conclude that the log10-transformed data better achieved the assumption of normality. Therefore, we would not report all the figures from Chapter 13.1: Figure 2, Figure 3, Figure 4, Figure 5, and Figure 6. I would report only Figure 2 — the raw data– with a few sentences: “The data were log10-transformed to address violations of normality assumption. Normality was assessed by calculation of the moment statistics skewness and kurtosis and Q-Q plots. All subsequent tests were conducted on the log10-transformed data set.” I wouldn’t report the exact results of the Q-Q plots or the moment statistics, but would include names or references to the specific tests used.
Note 2: Log10-transform would also likely improve unequal variances. Therefore, I would need to modify my sentences to account for this, eg, “The data were log10-transformed to address violations of normality and equal variance assumptions.”
Questions
Please skim read articles about statistical assumptions for common statistical tests:
- Choi (2005), click link to article.
- Hoekstra et al. (2012), click link to article.
- Olsen (2003), click link to article.
From the articles and your readings in Mike’s Biostatistics Book, answer the following questions
- What are some consequences if a researcher fails to check statistical assumptions?
- Explain why use of graphics techniques may be more important than results of statistical tests for checks of statistical assumptions.
- Briefly describe graphical and statistical tests of assumptions of (1) normality and (2) equal variances
- Pick a biological research journal for which you have online access to recent articles. Pick ten articles that used statistics (e.g., look for “t-test” or “ANOVA” or “regression”; exclude meta analysis and review articles — stick to primary research articles). Scan the Methods section and note whether or not you found a statement that confirms if the author(s) checked for violation of (1) normality and (2) equal variances. Construct a table.
- Review your results from question 3. Out of the ten articles, how many reported assumption checking? How does your result compare to those of Hoekstra et a; (2012)?
Quiz Chapter 13.2
Why tests of assumption are important
Chapter 13 contents
- Introduction
- ANOVA Assumptions
- Why tests of assumption are important
- Test assumption of normality
- Tests for Equal Variances
- References and suggested readings