18.1 – Multiple Linear Regression
Introduction
Last time we introduced simple linear regression:
- one independent X variable
- one dependent Y variable.
The linear relationship between Y and X was estimated by the method of Ordinary Least Squares (OLS). OLS minimizes the sum of squared distances between the observed responses, ![]() and responses predicted
and responses predicted ![]() by the line. Simple linear regression is analogous to our one-way ANOVA — one outcome or response variable and one factor or predictor variable (Chapter 12.2).
by the line. Simple linear regression is analogous to our one-way ANOVA — one outcome or response variable and one factor or predictor variable (Chapter 12.2).
But the world is complicated and so, our one-way ANOVA was extended to the more general case of two or more predictor (factor) variables (Chapter 14). As you might have guessed by now, we can extend simple regression to include more than one predictor variable. In fact, combining ANOVA and regression gives you the general linear model! And, you should not be surprised that statistics has extended this logic to include not only multiple predictor variables, but also multiple response variables. Multiple response variables falls into a category of statistics called multivariate statistics.
Like multi-way ANOVA, multiple regression is the extension of simple linear regression from one independent predictor variable to include two or more predictors. The benefit of this extension is obvious — our models gain realism. All else being equal, the more predictors, the better the model will be at describing and/or predicting the response. Things are not all equal, of course, and we’ll consider two complications of this basic premise, that more predictors are best; in some cases they are not.
However, before discussing the exceptions or even the complications of a multiple linear regression model, we begin by obtaining estimates of the full model, then introduce aspects of how to evaluate the model. We also introduce and whether a reduced model may be the preferred model.
R code
Multiple regression is easy to do in Rcmdr — recall that we used the general linear model function, lm(), to analyze one-way ANOVA and simple linear regression. In R Commander, we access lm() by
Rcmdr: Statistics → Fit model → Linear model
You may, however, access linear regression through R Commander
We use the same general linear model function for cases of multi-way ANOVA and for multiple regression problems. Simply enter more than one ratio-scale predictor variable and boom!
You now have yourself a multiple regression. You would then proceed to generate the ANOVA table for hypothesis testing
Rcmdr: Models → Hypothesis testing → ANOVA tables
From the output of the regression command, estimates of the coefficients along with standard errors for the estimate and results of t-tests for each coefficient against the respective null hypotheses for each coefficient are also provided. In our discussion of simple linear regression we introduced the components: the intercept, the slope, as well as the concept of model fit, as evidenced by R2, the coefficient of determination. These components exist for the multiple regression problem, too, but now we call the slopes partial regression slopes because there are more than one.
Our full multiple regression model becomes
![]()
where the coefficients β1, β2, … βn are the partial regression slopes and β0 is the Y-intercept for a model with 1 – n predictor variables. Each coefficient has a null hypothesis, each has a standard error, and therefore, each coefficient can be tested by t-test.
Now, regression, like ANOVA, is an enormous subject and we cannot do it justice in the few days we will devote to it. We can, however, walk you through a fairly typical example. I’ve posted a small data set diabetesCholStatin at the end of this page. Scroll down or click here. View the data set and complete your basic data exploration routine: make scatterplots and box plots. We think (predict) that body size and drug dose cause variation in serum cholesterol levels in adult men. But do both predict cholesterol levels?
Selecting the best model
We have two predictor variables, and we can start to see whether none, one, or both of the predictors contribute to differences in cholesterol levels. In this case, both contribute significantly. The power of multiple regression approaches is that it provides a simultaneous test of a model which may have many explanatory variables deemed appropriate to describe a particular response. More generally, it is sometimes advisable to think more philosophically about how to select a best model.
In model selection, some would invoke Occam’s razor — given a set of explanations, the simplest should be selected — to justify seeking simpler models. There are a number of approaches (forward selection, backward selection, or stepwise selection), and the whole effort of deciding among competing models is complicated with a number of different assumptions, strengths and weaknesses. I refer you to the discussion below, which of course is just a very brief introduction to a very large subject in (bio)statistics!
Let’s get the full regression model
The statistical model is
![]()
As written in R format, our model is ChLDL ~ BMI + Dose + Statin.
Note 1. BMI is ratio scale and Statin is categorical (two levels: Statin1, Statin2). Dose can be viewed as categorical, with five levels (5, 10, 20, 40, 80 mg), interval scale, or ratio scale. If we are make the assumption that the difference between 5, 10, up to 80 is meaningful, and that the effect of dose is at least proportional if not linear with respect to ChLDL, then we would treat Dose as ratio scale, not interval scale. That’s what we did here.
We can now proceed in R Commander to fit the model.
Rmdr: Statistics → Fit models → Linear model
How the model is inputted into linear model menu is shown in Figure 1.
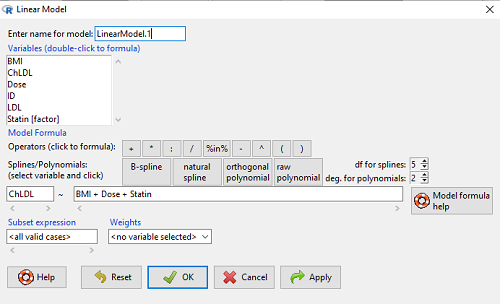
Figure 1. Screenshot of Rcmdr linear model menu with our model elements in place.
The output
summary(LinearModel.1)
Call:
lm(formula = ChLDL ~ BMI + Dose + Statin, data = cholStatins)
Residuals:
Min 1Q Median 3Q Max
-3.7756 -0.5147 -0.0449 0.5038 4.3821
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.016715 1.178430 0.863 0.39041
BMI 0.058078 0.047012 1.235 0.21970
Dose -0.014197 0.004829 -2.940 0.00411 **
Statin[Statin2] 0.514526 0.262127 1.963 0.05255 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.31 on 96 degrees of freedom
Multiple R-squared: 0.1231, Adjusted R-squared: 0.09565
F-statistic: 4.49 on 3 and 96 DF, p-value: 0.005407
Question. What are the estimates of the model coefficients (rounded)?
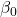 = intercept = 1.017
= intercept = 1.017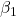 = slope for variable BMI = 0.058
= slope for variable BMI = 0.058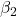 = slope for variable Dose = -0.014
= slope for variable Dose = -0.014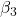 = slope for variable Statin = -0.515
= slope for variable Statin = -0.515
Question. Which of the three coefficients were statistically different from their null hypothesis (p-value < 5%)?
Answer: Only ![]() coefficient was judged statistically significant at the Type I error level of 5% (p = 0.0041). Of the four null hypotheses we have for the coefficients:
coefficient was judged statistically significant at the Type I error level of 5% (p = 0.0041). Of the four null hypotheses we have for the coefficients:
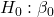 = 0
= 0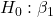 = 0
= 0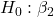 = 0; we only reject the null hypothesis for
= 0; we only reject the null hypothesis for Dosecoefficient.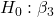 = 0)
= 0)
The important take-home concept is about the lack of a direct relationship between the magnitude of the estimate of the coefficient and the likelihood that it will be statistically significant! In absolute value terms ![]() , but
, but ![]() was not close to statistical significance (p = 0.220).
was not close to statistical significance (p = 0.220).
We generate a 2D scatterplot and include the regression lines (by group=Statin) to convey the relationship between at least one of the predictors (Fig 2).
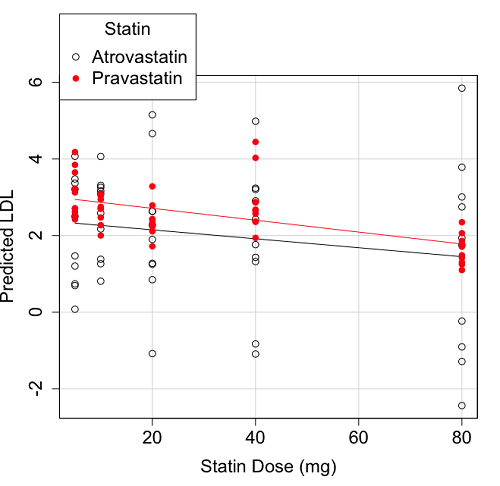
Figure 2. Scatter plot of predicted LDL against dose of a statin drug. Regression lines represent the different statin drugs (Statin1, Statin2).
Question. Based on the graph, can you explain why there will be no statistical differences between levels of the statin drug type, Statin1 (shown open circles) vs. Statin2 (shown closed red circles).
Because we have two predictors (BMI and Statin Dose), you may also elect to use a 3D-scatterplot. Here’s one possible result (Fig 3).
Figure 3. 3D plot of BMI and dose of Statin drugs on change in LDL levels (green Statin2, blue Statin1).
R code for Figure 3.
Graph made in Rcmdr: Graphs → 3D Graph → 3D scatterplot …
scatter3d(ChLDL~BMI+Dose|Statin, data=diabetesCholStatin, fit="linear", residuals=TRUE, parallel=FALSE, bg="white", axis.scales=TRUE, grid=TRUE, ellipsoid=FALSE)
Note 2: Figure 3 is a challenging graphic to interpret. I wouldn’t use it because it doesn’t convey a strong message. With some effort we can see the two planes representing mean differences between the two statin drugs across all predictors, but it’s a stretch. No doubt the graph can be improved by changing colors, for example, but I think the 2d plot Figure 2 works better). Alternatively, if the platform allows, you can use animation options to help your reader see the graph elements. Interactive graphics are very promising and, again, unsurprisingly, there are several R packages available. For this example, plot3d() of the package rgl can be used. Figure 4 is one possible version; I saved images and made animated gif.
Figure 4. An example of a possible interactive 3D plot; the file embedded in this page is not interactive, just an animation.
Diagnostic plots
While visualization concerns are important, let’s return to the statistics. All evaluations of regression equations should involve an inspection of the residuals. Inspection of the residuals allows you to decide if the regression fits the data; if the fit is adequate, you then proceed to evaluate the statistical significance of the coefficients.
The default diagnostic plots (Fig 5) R provides are available from
Rcmdr: Models → Graphs → Basic diagnostics plots
Four plots are returned
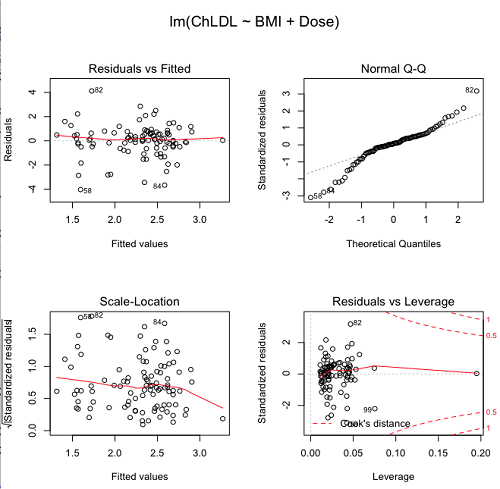
Figure 5. R’s default regression diagnostic plots.
Each of these diagnostic plots in Figure 4 gives you clues about the model fit.
- Plot of residuals vs. fitted helps you identify patterns in the residuals
- Normal Q-Q plot helps you to see if the residuals are approximately normally distributed
- Scale-location plot provides a view of the spread of the residuals
- The residuals vs. leverage plot allows you to identify influential data points.
We introduced these plots in Chapter 17.8 when we discussed fit of simple linear model to data. My conclusion? No obvious trend in residuals, so linear regression is a fit to the data; data not normally distributed Q-Q plot shows S-shape.
Interpreting the diagnostic plots for this problem
The “Normal Q-Q” plot allows us to view our residuals against a normal distribution (the dotted line). Our residuals do no show an ideal distribution: low for the first quartile, about on the line for intermediate values, then high for the 3rd and 4th quartile residuals. If the data were bivariate normal we would see the data fall along a straight line. The “S-shape” suggests log-transformation of the response and or one or more of the predictor variables.
There also seems to be a pattern in residuals vs the predicted (fitted) values. There is a trend of increasing residuals as cholesterol levels increase, which is particularly evident in the “scale-location” plot. Residuals tended to be positive at low and high doses, but negative at intermediate doses. This suggests that the relationship between predictors and cholesterol levels may not be linear, and it demonstrates what statisticians refer to as a monotonic spread of residuals.
The last diagnostic plot looks for individual points that influence, change, or “leverage” the regression — in other words, if a point is removed, does the general pattern change? If so, then the point had “leverage” and thus we need to decide whether or not to include the datum. diagnostic plots Cook’s distance is a measure of the influence of a point in regression. Points with large Cook’s distance values warrant additional checking.
The multicollinearity problem
Statistical model building is a balancing act by the statistician. While simpler models may be easier to interpret and, perhaps, to use, it is a basic truism that the more predictor variables the model includes, the more realistic the statistical model. However, each additional parameter that is added to the statistical model must be independent of all other parameters already in the model. To the extent that this assumption is violated, the problem is termed multicollinearity. If predictor variables are highly correlated, then they are essentially just linear combinations and do not provide independent evidence. For example, one would naturally not include two core body temperature variables in a statistical model on basal metabolic rate, one in degrees Fahrenheit and the other in degrees Celsius, because it is a simple linear conversion between the two units. This would be an example of structural collinearity: the collinearity is because of misspecification of the model variables. In contrast, collinearity among predictor variables may because the data are themselves correlated. For example, if multiple measures of body size are included: weight, height, length of arm, etc., then we would expect these to be correlated, ie, data multicollinearity.
Collinearity in statistical models may have a number of undesirable effects on a multiple regression model. These include
- estimates of coefficients not stable: with collinearity, values of coefficients depend on other variables in the model; if collinear predictors, then the assumption of independent predictor variables is violated.
- precision of the estimates decreases (standard error of estimates increase).
- statistical power decreases.
- p-values for individual coefficients not trust worthy.
Tolerance and Variance Inflation Factor
Absence of multicollinearity is important assumption of multiple regression. A partial test is to calculate product moment correlations among predictor variables. For example, when we calculate the correlation between BMI and Dose for our model, we get r = 0.101 (P = 0.3186), and therefore would tentatively conclude that there was little correlation between our predictor variables.
A number of diagnostic statistics have been developed to test for multicollinearity. Tolerance for a particular independent variable (Xi) is defined as 1 minus the proportion of variance it shares with the other independent variables in the regression analysis (1 − R2i) (O’Brien 2007). Tolerance reports the proportion of total variance explained by adding the Xi–th predictor variable that is unrelated to the other variables in the model. A small value for tolerance indicates multicollinearity — and that the predictor variable is nearly a perfect combination (linear) of the variables already in the model and therefore should be omitted from the model. Because tolerance is defined in relation to the coefficient of determination you can interpret a tolerance score as the unique variance accounted for by a predictor variable.
A second, related diagnostic of multicollinearity is called the Variance Inflation Factor, VIF. VIF is the inverse of tolerance.
![]()
VIF shows how much of the variance of a regression coefficient is increased because of collinearity with the other predictor variables in the model. VIF is easy to interpret: a tolerance of 0.01 has a VIF of 100; a tolerance of 0.1 has a VIF of 10; a tolerance of 0.5 has a VIF of 2, and so on. Thus, small values of tolerance and large values of VIF are taken as evidence of multicollinearity.
Rcmdr: Models → Numerical diagnostics → Variation-inflation factors
vif(RegModel.2) BMI Dose 1.010256 1.010256
A rule of thumb is that if VIF is greater than 5 then there is multicollinearity; with VIF values close to one we would conclude, like our results from the partial correlation estimate above, that there is little evidence for a problem of collinearity between the two predictor variables. They can therefore remain in the model.
Solutions for multicollinearity
If there is substantial multicollinearity then you cannot simply trust the estimates of the coefficients. Assuming that there hasn’t been some kind of coding error on your part, then you may need to find a solution. One solution is to drop one of the predictor variables and redo the regression model. Another option is to run what is called a Principal Components Regression (PCR). One takes the predictor variables and runs a Principal Component Analysis to reduce the number of variables, then the regression is run on the PCA components. By definition, the PCA components are independent of each other. Another option is to use ridge regression approach.
Note 3. Principal Component Analysis (PCA) is a dimensionality reduction technique that finds new uncorrelated variables (principal components) by maximizing variance. Principal Component Regression (PCR) uses PCA as a preprocessing step to create a regression model. The key difference is that PCA is an unsupervised method for data transformation, whereas PCR is a supervised regression technique that uses the components from PCA to predict an outcome. It’s probably not a great idea to use the acronym PCR in our biology reports; easily confused with polymerase chain reaction
Like any diagnostic rule, however, one should not blindly apply a rule of thumb. A VIF of 10 or more may indicate multicollinearity, but it does not necessarily lead to the conclusion that the linear regression model requires that the researcher reduce the number of predictor variables or analyze the problem using a different statistical method to address multicollinearity as the sole criteria of a poor statistical model. Rather, the researcher needs to address all of the other issues about model and parameter estimate stability, including sample size. Unless the collinearity is extreme (like a correlation of 1.0 between predictor variables!), larger sample sizes alone will work in favor of better model stability (by lowering the sample error) (O’Brien 2007).
Questions
- Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
- Please explain why the magnitude of the slope is not the key to statistical significance of a slope? Hint: look at the equation of the t-test for statistical significance of the slope.
- Consider the following scenario. A researcher measures his subjects multiple times for blood pressure over several weeks, then plots all of the values over time. In all, the data set consists of thousands of readings. He then proceeds to develop a model to explain blood pressure changes over time. What kind of collinearity is present in his data set? Explain your choice.
- We noted that
Dosecould be viewed as categorical variable. ConvertDoseto factor variable (fDose) and redo the linear model. Compare the summary output and discuss the additional coefficients.- Use Rcmdr: Data → Manage variables in active data set → Convert numeric Variables to Factors to create a new factor variable
fDose. It’s ok to use the numbers as factor levels.
- Use Rcmdr: Data → Manage variables in active data set → Convert numeric Variables to Factors to create a new factor variable
- We flagged the change in LDL as likely to be not normally distributed. Create a log10-transformed variable for
ChLDLand perform the multiple regression again.- Write the new statistical model
- Obtain the regression coefficients — are they statistically significant?
- Run basic diagnostic plots and evaluate for
Quiz Chapter 18.1
Multiple Linear Regression
Data set
| ID | Statin | Dose | BMI | LDL | ChLDL |
|---|---|---|---|---|---|
| 1 | Statin2 | 5 | 19.5 | 3.497 | 2.714778 |
| 2 | Statin1 | 20 | 20.2 | 4.268 | 1.276483 |
| 3 | Statin2 | 40 | 20.3 | 3.989 | 2.677377 |
| 4 | Statin2 | 20 | 20.3 | 3.502 | 2.430618 |
| 5 | Statin2 | 80 | 20.4 | 3.766 | 1.79463 |
| 6 | Statin2 | 20 | 20.6 | 3.44 | 2.234295 |
| 7 | Statin1 | 20 | 20.7 | 3.414 | 2.635305 |
| 8 | Statin1 | 10 | 20.8 | 3.222 | 0.809181 |
| 9 | Statin1 | 10 | 21.1 | 4.04 | 3.259599 |
| 10 | Statin1 | 40 | 21.2 | 4.429 | 1.763997 |
| 11 | Statin1 | 5 | 21.2 | 3.528 | 3.369377 |
| 12 | Statin1 | 40 | 21.5 | 3.01 | -0.827154 |
| 13 | Statin2 | 20 | 21.6 | 3.393 | 2.11172 |
| 14 | Statin1 | 10 | 21.7 | 4.512 | 3.166238 |
| 15 | Statin1 | 80 | 22 | 5.449 | 3.00833 |
| 16 | Statin2 | 10 | 22.2 | 4.03 | 3.05013 |
| 17 | Statin2 | 40 | 22.2 | 3.911 | 2.646034 |
| 18 | Statin2 | 10 | 22.2 | 3.724 | 2.945656 |
| 19 | Statin1 | 5 | 22.2 | 3.238 | 3.209584 |
| 20 | Statin2 | 10 | 22.5 | 4.123 | 3.088763 |
| 21 | Statin1 | 20 | 22.6 | 3.859 | 5.152548 |
| 22 | Statin1 | 10 | 23 | 4.926 | 2.58483 |
| 23 | Statin2 | 20 | 23 | 3.512 | 2.291975 |
| 24 | Statin1 | 5 | 23 | 3.838 | 1.469 |
| 25 | Statin2 | 20 | 23.1 | 3.548 | 2.34079 |
| 26 | Statin1 | 5 | 23.1 | 3.424 | 1.204346 |
| 27 | Statin1 | 40 | 23.2 | 3.709 | 3.238179 |
| 28 | Statin1 | 80 | 23.2 | 4.786 | 2.748643 |
| 29 | Statin1 | 20 | 23.3 | 4.103 | 1.250082 |
| 30 | Statin1 | 40 | 23.4 | 3.341 | 1.432292 |
| 31 | Statin1 | 10 | 23.5 | 3.828 | 1.381755 |
| 32 | Statin2 | 10 | 23.8 | 4.02 | 3.039187 |
| 33 | Statin1 | 20 | 23.8 | 3.942 | 0.848328 |
| 34 | Statin2 | 20 | 23.8 | 2.89 | 1.721163 |
| 35 | Statin1 | 80 | 23.9 | 3.326 | 1.939346 |
| 36 | Statin1 | 10 | 24.1 | 4.071 | 3.090741 |
| 37 | Statin1 | 40 | 24.1 | 4.222 | 1.322305 |
| 38 | Statin2 | 10 | 24.1 | 3.44 | 2.472223 |
| 39 | Statin1 | 5 | 24.2 | 3.507 | 0.076817 |
| 40 | Statin2 | 20 | 24.2 | 3.647 | 2.425758 |
| 41 | Statin2 | 80 | 24.3 | 3.812 | 1.710575 |
| 42 | Statin2 | 40 | 24.3 | 3.305 | 1.940572 |
| 43 | Statin2 | 5 | 24.3 | 3.455 | 2.502214 |
| 44 | Statin2 | 5 | 24.4 | 4.258 | 3.228008 |
| 45 | Statin1 | 5 | 24.4 | 4.16 | 3.477747 |
| 46 | Statin2 | 80 | 24.4 | 4.128 | 2.063247 |
| 47 | Statin1 | 80 | 24.5 | 4.507 | 3.784422 |
| 48 | Statin1 | 5 | 24.5 | 3.553 | 0.695709 |
| 49 | Statin2 | 10 | 24.5 | 3.616 | 2.69987 |
| 50 | Statin2 | 80 | 24.6 | 3.372 | 1.300401 |
| 51 | Statin2 | 80 | 24.6 | 3.667 | 1.418109 |
| 52 | Statin2 | 5 | 24.7 | 3.854 | 3.126671 |
| 53 | Statin1 | 80 | 24.7 | 3.32 | -1.286439 |
| 54 | Statin2 | 5 | 24.7 | 3.756 | 2.423664 |
| 55 | Statin1 | 40 | 24.8 | 4.398 | 2.907473 |
| 56 | Statin2 | 40 | 24.9 | 3.621 | 2.362429 |
| 57 | Statin1 | 10 | 25 | 3.17 | 1.264656 |
| 58 | Statin1 | 80 | 25.1 | 3.424 | -2.436908 |
| 59 | Statin2 | 10 | 25.1 | 3.196 | 2.001465 |
| 60 | Statin2 | 80 | 25.2 | 3.367 | 1.100704 |
| 61 | Statin1 | 80 | 25.2 | 3.067 | -0.23154 |
| 62 | Statin1 | 20 | 25.3 | 3.678 | 4.662866 |
| 63 | Statin2 | 5 | 25.5 | 4.077 | 2.611705 |
| 64 | Statin1 | 20 | 25.5 | 3.678 | 2.633053 |
| 65 | Statin2 | 5 | 25.6 | 4.994 | 4.180082 |
| 66 | Statin1 | 20 | 25.8 | 3.699 | 1.899031 |
| 67 | Statin1 | 10 | 25.9 | 3.507 | 4.063757 |
| 68 | Statin2 | 20 | 25.9 | 3.445 | 2.303761 |
| 69 | Statin1 | 5 | 26 | 4.025 | 2.501427 |
| 70 | Statin1 | 5 | 26.3 | 3.616 | 0.740863 |
| 71 | Statin2 | 40 | 26.4 | 3.937 | 2.573321 |
| 72 | Statin2 | 40 | 26.4 | 3.823 | 2.363839 |
| 73 | Statin1 | 10 | 26.7 | 4.46 | 2.174198 |
| 74 | Statin2 | 5 | 26.7 | 5.03 | 3.845271 |
| 75 | Statin2 | 10 | 26.7 | 3.73 | 2.708896 |
| 76 | Statin2 | 10 | 26.7 | 3.232 | 2.272627 |
| 77 | Statin1 | 80 | 26.8 | 3.693 | 1.751169 |
| 78 | Statin2 | 80 | 27 | 4.108 | 1.86131 |
| 79 | Statin2 | 40 | 27.2 | 5.398 | 4.028977 |
| 80 | Statin2 | 80 | 27.2 | 4.517 | 2.348903 |
| 81 | Statin2 | 20 | 27.3 | 3.901 | 2.790047 |
| 82 | Statin1 | 80 | 27.3 | 5.247 | 5.848545 |
| 83 | Statin2 | 80 | 27.4 | 3.507 | 1.247863 |
| 84 | Statin1 | 20 | 27.4 | 3.807 | -1.079928 |
| 85 | Statin2 | 80 | 27.6 | 3.574 | 1.486789 |
| 86 | Statin1 | 40 | 27.8 | 4.16 | 2.427753 |
| 87 | Statin2 | 20 | 28 | 4.501 | 3.284648 |
| 88 | Statin2 | 5 | 28.1 | 3.621 | 2.699007 |
| 89 | Statin1 | 40 | 28.2 | 3.652 | -1.091256 |
| 90 | Statin2 | 40 | 28.2 | 4.191 | 2.874231 |
| 91 | Statin2 | 40 | 28.4 | 5.791 | 4.445454 |
| 92 | Statin1 | 40 | 28.6 | 4.698 | 3.202874 |
| 93 | Statin1 | 5 | 29 | 4.32 | 4.070753 |
| 94 | Statin2 | 10 | 29.1 | 3.776 | 2.751281 |
| 95 | Statin2 | 5 | 29.2 | 4.703 | 3.64949 |
| 96 | Statin2 | 40 | 29.9 | 4.128 | 2.864691 |
| 97 | Statin1 | 40 | 30.4 | 4.693 | 4.983704 |
| 98 | Statin1 | 20 | 30.4 | 4.123 | 2.273898 |
| 99 | Statin1 | 80 | 30.5 | 3.921 | -0.903438 |
| 100 | Statin1 | 10 | 36.5 | 4.175 | 3.311437 |
Chapter 18 contents
17.8 – Assumptions and model diagnostics for Simple Linear Regression
Introduction
The assumptions for all linear regression:
- Linear model is appropriate.
The data are well described (fit) by a linear model. - Independent values of Y and equal variances.
Although there can be more than one Y for any value of X, the Y‘s cannot be related to each other (that’s what we mean by independent). Since we allow for multiple Y‘s for each X, then we assume that the variances of the range of Y‘s are equal for each X value (this is similar to our ANOVA assumptions for equal variance by groups). Another term for equal variances is homoscedasticity. - Normality.
For each X value there is a normal distribution of Y‘s (think of doing the experiment over and over). - Error
The residuals (error) are normally distributed with a mean of zero.
Note the mnemonic device: Linear, Independent, Normal, Error or LINE.
Each of the four elements will be discussed below in the context of Model Diagnostics. These assumptions apply to how the model fits the data. There are other assumptions that, if violated, imply you should use a different method for estimating the parameters of the model.
Ordinary least squares makes the additional assumption about the quality of the independent variable that e that measurement of X is done without error. Measurement error is a fact of life in science, but the influence of error on regression differs if the error is associated with the dependent or independent variable. Measurement error in the dependent variable increases the dispersion of the residuals but will not affect the estimates of the coefficients; error associated with the independent variables, however, will affect estimates of the slope. In short, error in X leads to biased estimates of the slope.
The equivalent, but less restrictive practical application of this assumption is that the error in X is at least negligible compared to the measurements in the dependent variable.
Multiple regression makes one more assumption, about the relationship between the predictor variables (the X variables). The assumption is that there is no multicollinearity, a subject we will bring up next time (see Chapter 18).
Model diagnostics
We just reviewed how to evaluate the estimates of the coefficients of the model. Now we need to address a deeper meaning — how well the model explains the data. Consider a simple linear regression first. If Ho: b = 0 is not rejected — then the slope of the regression equation is taken to not differ from zero. We would conclude that if repeated samples were drawn from the population, on average, the regression equation would not fit the data well (lots of scatter) and it would not yield useful prediction.
However, recall that we assume that the fit is linear. One assumption we make in regression is that a line can, in fact, be used to describe the relationship between X and Y.
Here are two very different situations where the slope = 0.
Example 1. Linear Slope = 0, No relationship between X and Y
Example 2. Linear Slope = 0, A significant relationship between X and Y
But even if Ho: b = 0 is rejected (and we conclude that a linear relationship between X and Y is present), we still need to be concerned about the fit of the line to the data — the relationship may be more nonlinear than linear, for example. Here are two very different situations where the slope is not equal to 0.
Example 3. Linear Slope > 0, a linear relationship between X and Y
Example 4. Linear Slope > 0, curve-linear relationship between X and Y
How can you tell the difference? There are many regression diagnostic tests, many more than we can cover, but you can start with looking at the coefficient of determination (low R2 means low fit to the line), and we can look at the pattern of residuals plotted against the either the predicted values or the X variables (my favorite). The important points are:
- In linear regression, you fit a model (the slope + intercept) to the data;
- We want the usual hypothesis tests (are the coefficients different from zero?) and
- We need to check to see if the model fits the data well. Just like in our discussions of chi-square, a “perfect fit would mean that the difference between our model and the data would be zero.
Graph options
Using residual plots to diagnose regression equations
Yes, we need to test the coefficients (intercept Ho = 0; slope Ho = 0) of a regression equation, but we also must decide if a regression is an appropriate description of the data. This topic includes the use of diagnostic tests in regression. We address this question chiefly by looking at
- scatterplots of the independent (predictor) variable(s) vs. dependent (response) variable(s).
what patterns appear between X and Y? Do your eyes tell you “Line”? “Curve”? “No relation”? - coefficient of determination
closer to zero than to one? - patterns of residuals plotted against the X variables (other types of residual plots are used to, this is one of my favorites)
Our approach is to utilize graphics along with statistical tests designed to address the assumptions.
One typical choice is to see if there are patterns in the residual values plotted against the predictor variable. If the LINE assumptions hold for your data set, then the residuals should have a mean of zero with scatter about the mean. Deviations from LINE assumptions will show up in residual plots.
Here are examples of POSSIBLE outcomes:
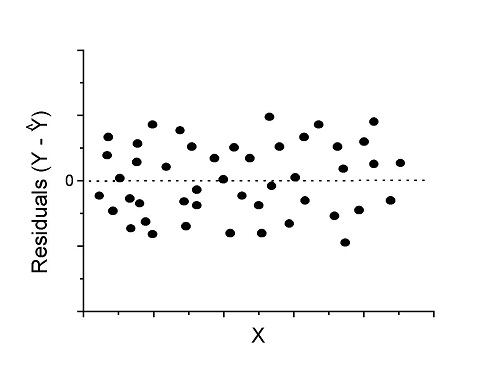
Figure 1. An ideal plot of residuals
Solution: Proceed! Assumptions of linear regression met.
Compare to plots of residuals that differ from the ideal.
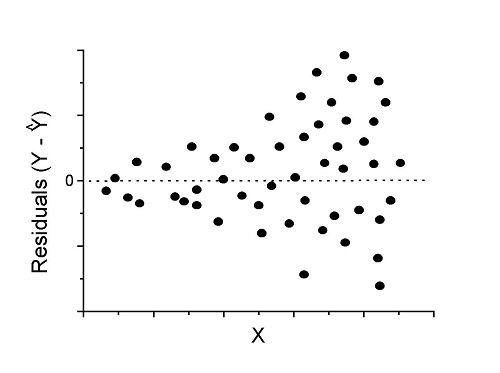
Figure 2. We have a problem. Residual plot shows a “funnel” shape — unequal variance (aka heteroscedasticity).
Solution. Try a transform like the log10-transform.
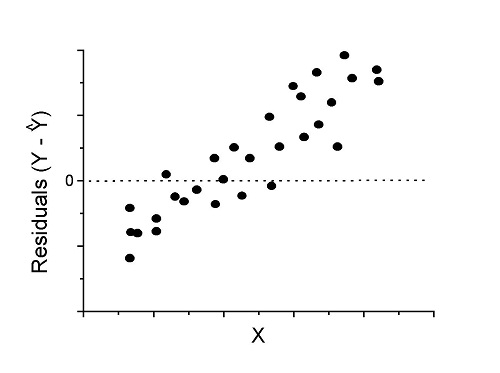
Figure 3. Problem. Residual plot shows systematic trend.
Solution. Linear model a poor fit; May be related to measurement errors for one or more predictor variables. Try adding an additional predictor variable or model the error in your general linear model.
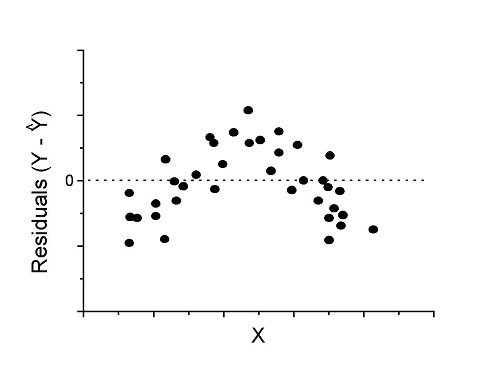
Figure 4. Problem. Residual plot shows nonlinear trend.
Solution. Transform data or use more complex model
This is a good time to mention that in statistical analyses, one often needs to do multiple rounds of analyses, involving description and plots, tests of assumptions, tests of inference. With regression, in particular, we also need to decide if our model (e.g., linear equation) is a good description of the data.
Diagnostic plot examples
Return to our example.Tadpole dataset. To obtain residual plots, Rcmdr: Models → Graphs → Basic diagnostic plots yields four graphs. Graph A is
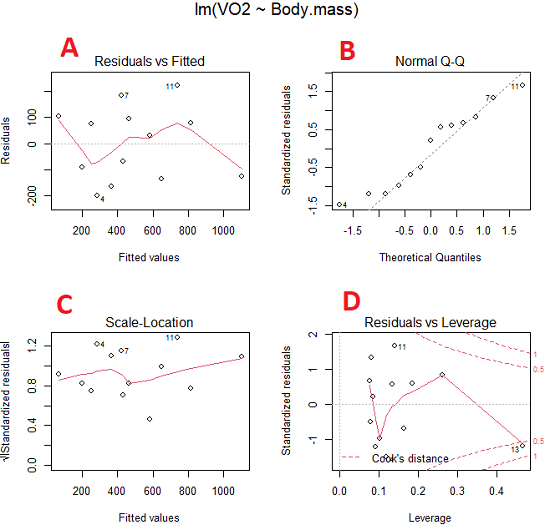
Figure 5. Basic diagnostic plots. A: residual plot; B: Q-Q plot of residuals; C: Scale-location (aka spread-location) plot; D: leverage residual plot.
In brief, we look at plots
A, the residual plot, to see if there are trends in the residuals. We are looking for a spread of points equally above and below the mean of zero. In Figure 5 we count seven points above and six points below zero so there’s no indication of a trend in the residuals vs the fitted VO2 (Y) values.
B, the Q-Q plot is used to see if normality holds. As discussed before, if our data are more or less normally distributed, then points will fall along a straight line in a Q-Q plot.
C, the Scale- or spread-location plot is used to verify equal variances of errors.
D, Leverage plot — looks to see if an outlier has leverage on the fit of the line to the data, i.e., changes the slope. Additionally, provides location of Cook’s distance measure (dashed red lines). Cook’s distance measures the effect on the regression by removing one point at a time and then fitting a line to the data. Points outside the dashed lines have influence.
A note of caution about over-thinking with these plots. R provides a red line to track the points. However, these lines are guides, not judges. We humans are generally good at detecting patterns, but with data visualization, there is the risk of seeing patterns where none exits. In particular, recognizing randomness is not easy. If anything, we may tend to see patterns where none exist, termed apophenia. So yes, by all means look at the graphs, but do so with a plan: red line more or less horizontal? Then no pattern and the regression model is a good fit to the data.
Statistical test options
After building linear models, run statistical diagnostic tests that compliment graphics approaches. Here, simply to introduce the tests, I included the Gosner factor as potential predictor — many of these numerical diagnostics are more appropriate for models with more than one predictor variable, the subject of the next chapter.
Results for the model ![]() were:
were:
lm(formula = VO2 ~ Body.mass + Gosner, data = frogs)
Residuals:
Min 1Q Median 3Q Max
-163.12 -125.53 -20.27 83.71 228.56
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -595.37 239.87 -2.482 0.03487 *
Body.mass 431.20 115.15 3.745 0.00459 **
Gosner[T.II] 64.96 132.83 0.489 0.63648
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 153.4 on 9 degrees of freedom
(1 observation deleted due to missingness)
Multiple R-squared: 0.8047, Adjusted R-squared: 0.7613
F-statistic: 18.54 on 2 and 9 DF, p-value: 0.0006432
Question. Note the p-value for the coefficients — do we include a model with both body mass and Gosner Stage? See Chapter 18.5 – Selecting the best model
In R Commander, the diagnostic test options are available via
Rcmdr: Models → Numerical diagnostics
Variance inflation factors (VIF): used to detect multicollinearity among the predictor variables. If correlations are present among the predictor variables, then you can’t rely on the the coefficient estimates — whether predictor A causes change in the response variable depends on whether the correlated B predictor is also included in the model. If correlation between predictor A and B, the statistical effect is increased variance associated with the error of the coefficient estimates. There are VIF for each predictor variable. A VIF of one means there is no correlation between that predictor and the other predictor variables. A VIF of 10 is taken as evidence of serious multicollinearity in the model.
R code:
vif(RegModel.2)
R output
Body.mass Gosner
2.186347 2.186347
round(cov2cor(vcov(RegModel.2)), 3) # Correlations of parameter estimates
(Intercept) Body.mass Gosner[T.II]
(Intercept) 1.000 -0.958 0.558
Body.mass -0.958 1.000 -0.737
Gosner[T.II] 0.558 -0.737 1.000
Our interpretation
The output shows a matrix of correlations; the correlation between the two predictor variables in this model was 0.558, large per our discussions in Chapter 16.1 . Not really surprising as older tadpoles (Gosner stage II), are larger tadpoles. We’ll leave the issue of multicollinearity to Chapter 18.1.
Breusch-Pagan test for heteroscedasticity… Recall that heteroscedasticity is another name for unequal variances. This tests pertains to the residuals; we used the scale (spread) location plot (C) above. The test statistic can be calculated as ![]() . The function call is part of the car package, which we download as part of the Rcmdr package. R Commander will pop up a menu of choices for this test. For starters, accept the defaults, enter nothing, and click OK.
. The function call is part of the car package, which we download as part of the Rcmdr package. R Commander will pop up a menu of choices for this test. For starters, accept the defaults, enter nothing, and click OK.
R code:
library(zoo, pos=16) library(lmtest, pos=16) bptest(VO2 ~ Body.mass + Gosner, varformula = ~ fitted.values(RegModel.2), studentize=FALSE, data=frogs)
R output
Breusch-Pagan test data: VO2 ~ Body.mass + Gosner BP = 0.0040766, df = 1, p-value = 0.9491
Our interpretation
A p-value less than 5% would be interpreted as statistically significant unequal variances among the residuals.
Durbin-Watson for autocorrelation… used to detect serial (auto)correlation among the residuals, specifically, adjacent residuals. Substantial autocorrelation can mean Type I error for one or more model coefficients. The test returns a test statistic that ranges from 0 to 4. A value at midpoint (2) suggests no autocorrelation. Default test for autocorrelation is r > 0.
R code:
dwtest(VO2 ~ Body.mass + Gosner, alternative="greater", data=frogs)
R output
Durbin-Watson test data: VO2 ~ Body.mass + Gosner DW = 2.1336, p-value = 0.4657 alternative hypothesis: true autocorrelation is greater than 0
Our interpretation
The DW statistic was about 2. Per our definition, we conclude no correlations among the residuals.
RESET test for nonlinearity… The Ramsey Regression Equation Specification Error Test is used to interpret whether or not the equation is properly specified. The idea is that fit of the equation is compared against alternative equations that account for possible nonlinear dependencies among predictor and dependent variables. The test statistic follows an F distribution.
R code:
resettest(VO2 ~ Body.mass + Gosner, power=2:3, type="regressor", data=frogs)
R output
RESET test data: VO2 ~ Body.mass + Gosner RESET = 0.93401, df1 = 2, df2 = 7, p-value = 0.437
Our interpretation
Also a nonsignificant test — adding a power term to account for a nonlinear trend gained no model fit benefits, p-value much greater than 5%.
Bonferroni outlier test. Reports the Bonferroni p-values for testing each observation to see if it can be considered to be a mean-shift outlier (a method for identifying potential outlier values — each value is in turn replaced with the mean of its nearest neighbors, Lehhman et al 2020). Compare to results of VIF. The objective of outlier and VIF tests is to verify that the regression model is stable, insensitive to potential leverage (outlier) values.
R code:
outlierTest(LinearModel.3)
R output
No Studentized residuals with Bonferroni p < 0.05 Largest |rstudent|: rstudent unadjusted p-value Bonferroni p 7 1.958858 0.085809 NA
Our interpretation
We ran the outlierTest() function on LinearModel.3 to check for influential outliers using Studentized residuals with Bonferroni correction.
The output shows “No Studentized residuals with Bonferroni p < 0.05”, which means no data points were statistically flagged as outliers after adjusting for multiple comparisons. The largest Studentized residual was 1.96 (point 7), with an unadjusted p-value of 0.086. Although this is the largest deviation from the regression line, it is not statistically significant. Therefore, we conclude that no single observation is exerting undue influence on the model fit, and we can be confident that our regression results are not driven by outliers.
Response transformation… The concept of transformations of data to improve adherence to statistical assumptions was introduced in Chapters 3.2, 13.1, and 15.
R code:
summary(powerTransform(LinearModel.3, family="bcPower"))
R output
bcPower Transformation to Normality
Est Power Rounded Pwr Wald Lwr Bnd Wald Upr Bnd
Y1 0.9987 1 0.2755 1.7218
Likelihood ratio test that transformation parameter is equal to 0
(log transformation)
LRT df pval
LR test, lambda = (0) 7.374316 1 0.0066162
Likelihood ratio test that no transformation is needed
LRT df pval
LR test, lambda = (1) 0.00001289284 1 0.99714
Our interpretation
The Box-Cox analysis suggests that the response variable for LinearModel.3 can remain on its original scale. No power transformation was needed, and any standard linear regression inference is appropriate. The estimated transformation parameter was close to one, which suggests that the original response variable did not need a transformation. The LRT tests whether a log transformation (lambda = 0) would improve normality. The significant p-value, 0.0066, for the LRT indicates that a log transformation would not be the best choice, consistent with lambda ≈ 1. And last, the very high p-value, 0.99714, confirms that no transformation is necessary; the model residuals were close to normal.
Questions
- Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
- Referring to Figures 1 – 4 on this page, which plot best suggests a regression line fits the data?
- Return to the electoral college data set and your linear models of Electoral vs. POP_2010 and POP_2019. Obtain the four basic diagnostic plots and comment on the fit of the regression line to the electoral college data.
- residual plot
- Q-Q plot
- Scale-location plot
- Leverage plot
- With respect to your answers in question 2, how well does the electoral college system reflect the principle of one person one vote?
Quiz Chapter 17.8
Assumptions and model diagnostics for Simple Linear Regression
Chapter 17 contents
- Introduction
- Simple Linear Regression
- Relationship between the slope and the correlation
- Estimation of linear regression coefficients
- OLS, RMA, and smoothing functions
- Testing regression coefficients
- Regression model fit
- Assumptions and model diagnostics for Simple Linear Regression
- References and suggested readings (Ch17 & 18)
17.7 – Regression model fit
Introduction
In Chapter 17.5 and 17.6 we introduced the example of tadpoles body size and oxygen consumption. We ran a simple linear regression, with the following output from R
RegModel.1 <- lm(VO2~Body.mass, data=example.Tadpole)
summary(RegModel.1)
Call:
lm(formula = VO2 ~ Body.mass, data = example.Tadpole)
Residuals:
Min 1Q Median 3Q Max
-202.26 -126.35 30.20 94.01 222.55
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -583.05 163.97 -3.556 0.00451 **
Body.mass 444.95 65.89 6.753 0.0000314 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 145.3 on 11 degrees of freedom
Multiple R-squared: 0.8057, Adjusted R-squared: 0.788
F-statistic: 45.61 on 1 and 11 DF, p-value: 0.00003144
You should be able to pick out the estimates of slope and intercept from the table (intercept was -583 and slope was 445). Additionally, as part of your interpretation of the model, you should be able to report how much variation in VO2 was explained by tadpole body mass (coefficient of determination, ![]() , was 0.81, which means about 81% of variation in oxygen consumption by tadpoles is explained by knowing the body mass of the tadpole.
, was 0.81, which means about 81% of variation in oxygen consumption by tadpoles is explained by knowing the body mass of the tadpole.
What’s left to do? We need to evaluate how well our model fits the data, i.e., we evaluate regression model fit. By fit we mean, how well does our model agree to the raw data? A poor fit and the model predictions are far from the raw data; a good fit, the model accurately predicts the raw data. In other words, if we draw a line through the raw data, a good fit line crosses through most of the points — a poor fit the line rarely passes through the cloud of points.
How to judge fit? This we can do by evaluating the error components relative to the portion of the model that explains the data. Additionally, we can perform a number of diagnostics of the model relative to the assumptions we made to perform linear regression. These diagnostics form the subject of Chapter 17.8. Here, we ask how well does the model,
![]()
fit the data?
Model fit statistics
The second part of fitting a model is to report how well the model fits the data. The next sections apply to this aspect of model fitting. The first area to focus on is the magnitude of the residuals: the greater the spread of residuals, the less well a fitted line explains the data.
In addition to the output from lm() function, which focuses on the coefficients, we typically generate the ANOVA table also.
Anova(RegModel.1, type="II")
Anova Table (Type II tests)
Response: VO2
Sum Sq Df F value Pr(>F)
Body.mass 962870 1 45.605 0.00003144 ***
Residuals 232245 11
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Standard error of regression
S, the Residual Standard Error (aka Standard error of regression), is an overall measure to indicate the accuracy of the fitted line: it tells us how good the regression is in predicting the dependence of response variable on the independent variable. A large value for S indicates a poor fit. One equation for S is given by
![]()
In the above example, S = 145.3 (underlined, bold in regression output above). We can see how if ![]() is large, S will be large indicating poor fit of the linear model to the data. However, by itself S is not of much value as a diagnostic as it is difficult to know what to make of 145.3, for example. Is this a large value for S? Is it small? We don’t have any context to judge S, so additional diagnostics have been developed.
is large, S will be large indicating poor fit of the linear model to the data. However, by itself S is not of much value as a diagnostic as it is difficult to know what to make of 145.3, for example. Is this a large value for S? Is it small? We don’t have any context to judge S, so additional diagnostics have been developed.
Coefficient of determination
![]() , the coefficient of determination, is also used to describe model fit.
, the coefficient of determination, is also used to describe model fit. ![]() can take on values between 0 and 1 (0% to 100%). A good model fit has a high
can take on values between 0 and 1 (0% to 100%). A good model fit has a high ![]() value. In our example above,
value. In our example above, ![]() or 80.57%. One equation for
or 80.57%. One equation for ![]() is given by
is given by
![]()
A value of ![]() close to 1 means that the regression “explains” nearly all of the variation in the response variable, and would indicate the model is a good fit to the data. As we mentioned in Chapter 17.2, the coefficient of determination,
close to 1 means that the regression “explains” nearly all of the variation in the response variable, and would indicate the model is a good fit to the data. As we mentioned in Chapter 17.2, the coefficient of determination, ![]() , is the squared value of r, the product moment correlation.
, is the squared value of r, the product moment correlation.
Adjusted R-squared
Before moving on we need to remark on the difference between ![]() and adjusted
and adjusted ![]() . For Simple Linear Regression there is but one predictor variable, X; for multiple regression there can be many additional predictor variables. Without some correction,
. For Simple Linear Regression there is but one predictor variable, X; for multiple regression there can be many additional predictor variables. Without some correction, ![]() will increase with each additional predictor variables. This doesn’t mean the model is more useful, however, and in particular, one cannot compare
will increase with each additional predictor variables. This doesn’t mean the model is more useful, however, and in particular, one cannot compare ![]() between models with different numbers of predictors. Therefore, an adjustment is used so that the coefficient of determination remains a useful way to assess how reliable a model is and to permit comparisons of models. Thus, we have the Adjusted
between models with different numbers of predictors. Therefore, an adjustment is used so that the coefficient of determination remains a useful way to assess how reliable a model is and to permit comparisons of models. Thus, we have the Adjusted ![]() , which is calculated as
, which is calculated as
![]()
In our example above, Adjusted R 2 = 0.3806 or 38.06%.
Which should you report? Adjusted ![]() ,because it is independent of the number of parameters in the model.
,because it is independent of the number of parameters in the model.
Both ![]() and S are useful for regression diagnostics, a topic which we will discuss next (Chapter 17.8).
and S are useful for regression diagnostics, a topic which we will discuss next (Chapter 17.8).
Questions
- Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
- As you may know, the US census is used to reapportion congress and also to determine the number of electoral college votes. In honor of the election for US President that’s just days away, in the next series of questions in this Chapter and subsequent sections of Chapter 17 and 18, I’ll ask you to conduct a regression analysis on the electoral college. For starters, make the regression of Electoral votes on 2010 census population. (Ignore for now the other columns, just focus on POP_2019 and Electoral.) Report the
- regression coefficients (slope, intercept)
- percent of the variation in electoral college votes explained by the regression (
 ).
).
- Make a scatterplot and add the regression line to the plot
Quiz Chapter 17.7
Regression model fit
Data set
USA population year 2010 and year 2019 with Electoral College counts
| State | Region | Division | POP_2010 | POP_2019 | Electoral |
|---|---|---|---|---|---|
| Alabama | South | East South Central | 4779736 | 4903185 | 9 |
| Alaska | West | Pacific | 710231 | 731545 | 3 |
| Arizona | West | Mountain | 6392017 | 7278717 | 11 |
| Arkansas | South | West South Central | 2915918 | 3017804 | 6 |
| California | West | Pacific | 37253956 | 39512223 | 55 |
| Colorado | West | Mountain | 5029196 | 5758736 | 9 |
| Connecticut | Northeast | New England | 3574097 | 3565287 | 7 |
| Delaware | South | South Atlantic | 897934 | 982895 | 3 |
| District of Columbia | South | South Atlantic | 601723 | 705749 | 3 |
| Florida | South | South Atlantic | 18801310 | 21477737 | 29 |
| Georgia | South | South Atlantic | 9687653 | 10617423 | 16 |
| Hawaii | West | Pacific | 1360301 | 1415872 | 4 |
| Idaho | West | Mountain | 1567582 | 1787065 | 4 |
| Illinois | Midwest | East North Central | 12830632 | 12671821 | 20 |
| Indiana | Midwest | East North Central | 6483802 | 6732219 | 11 |
| Iowa | Midwest | West North Central | 3046355 | 3155070 | 6 |
| Kansas | Midwest | West North Central | 2853118 | 2913314 | 6 |
| Kentucky | South | East South Central | 4339367 | 4467673 | 8 |
| Louisiana | South | West South Central | 4533372 | 4648794 | 8 |
| Maine | Northeast | New England | 1328361 | 1344212 | 4 |
| Maryland | South | South Atlantic | 5773552 | 6045680 | 10 |
| Massachusetts | Northeast | New England | 6547629 | 6892503 | 11 |
| Michigan | Midwest | East North Central | 9883640 | 9883635 | 16 |
| Minnesota | Midwest | West North Central | 5303925 | 5639632 | 10 |
| Mississippi | South | East South Central | 2967297 | 2976149 | 6 |
| Missouri | Midwest | West North Central | 5988927 | 6137428 | 10 |
| Montana | West | Mountain | 989415 | 1068778 | 3 |
| Nebraska | Midwest | West North Central | 1826341 | 1934408 | 5 |
| Nevada | West | Mountain | 2700551 | 3080156 | 6 |
| New Hampshire | Northeast | New England | 1316470 | 1359711 | 4 |
| New Jersey | Northeast | Mid-Atlantic | 8791894 | 8882190 | 14 |
| New Mexico | West | Mountain | 2059179 | 2096829 | 5 |
| New York | Northeast | Mid-Atlantic | 19378102 | 19453561 | 29 |
| North Carolina | South | South Atlantic | 9535483 | 10488084 | 15 |
| North Dakota | Midwest | West North Central | 672591 | 762062 | 3 |
| Ohio | Midwest | East North Central | 11536504 | 11689100 | 18 |
| Oklahoma | South | West South Central | 3751351 | 3956971 | 7 |
| Oregon | West | Pacific | 3831074 | 4217737 | 7 |
| Pennsylvania | Northeast | Mid-Atlantic | 12702379 | 12801989 | 20 |
| Rhode Island | Northeast | New-England | 1052567 | 1059361 | 4 |
| South Carolina | South | South-Atlantic | 4625364 | 5148714 | 9 |
| South Dakota | Midwest | West-North-Central | 814180 | 884659 | 3 |
| Tennessee | South | East-South-Central | 6346105 | 6829174 | 11 |
| Texas | South | West-South-Central | 25145561 | 28995881 | 38 |
| Utah | West | Mountain | 2763885 | 3205958 | 6 |
| Vermont | Northeast | New-England | 625741 | 623989 | 3 |
| Virginia | South | South-Atlantic | 8001024 | 8535519 | 13 |
| Washington | West | Pacific | 6724540 | 7614893 | 12 |
| West Virginia | South | South-Atlantic | 1852994 | 1792147 | 5 |
| Wisconsin | Midwest | East-North-Central | 5686986 | 5822434 | 10 |
| Wyoming | West | Mountain | 563626 | 578759 | 3 |
Chapter 17 contents
- Introduction
- Simple Linear Regression
- Relationship between the slope and the correlation
- Estimation of linear regression coefficients
- OLS, RMA, and smoothing functions
- Testing regression coefficients
- ANCOVA – Analysis of covariance
- Regression model fit
- Assumptions and model diagnostics for Simple Linear Regression
- References and suggested readings (Ch17 & 18)
17.5 – Testing regression coefficients
Introduction
Whether the goal is to create a predictive model or an explanatory model, then there are two related questions the analyst asks about the linear regression model fitted to the data:
- Does a line actually fit the data
- Is the linear regression statistically significant?
We will turn to the first question pertaining to fit in time, but for now, focus on the second question.
Like the one-way ANOVA, we have the null and alternative hypothesis for the regression model itself. We write our hypotheses for the regression
H0 : linear regression fits the data vs. HA : linear regression does not fit
We see from the output in R and Rcmdr that an ANOVA table has been provided. Thus, the test of the regression is analogous to an ANOVA — we partition the overall variability in the response variable into two parts: the first part is the part of the variation that can be explained by there being a linear regression (the linear regression sum of squares) plus a second part that accounts for the rest of the variation in the data that is not explained by the regression (the residual or error sum of squares). Thus, we have
![]()
As we did in ANOVA, we calculate Mean Squares ![]() , where x refers to either “regression” or “residual” sums of squares and degrees of freedom. We then calculate F-values, the test statistics for the regression, to test the null hypothesis.
, where x refers to either “regression” or “residual” sums of squares and degrees of freedom. We then calculate F-values, the test statistics for the regression, to test the null hypothesis.
The degrees of freedom (DF) in simple linear regression are always
![]()
where
![]()
and FDFregression, DFresidual are compared to the critical value at Type I error rate α, with DFregression, DFresidual.
Linear regression inference
Estimation of the slope and intercept is a first step and should be accompanied by the calculation of confidence intervals.
What we need to know if we are to conclude that there’s a functional relationship between the X and Y variable is whether the same relationship exists in the population. We’ve sampled from the population, calculated an equation to describe the relationship between them. However, just as in all cases of inferential statistics, we need to consider the possibility that, through chance alone, we may have committed a Type I error.
The graph below (Fig 1) shows a possible outcome under a scenario in which the statistical analyst would likely conclude that there is a statistically significant linear model fit to the data, but the true relationship in the population was a slope of zero. How can this happen? Under this scenario, by chance alone the researchers sampled points (circled in red) from the population that fall along a line. We will conclude that there is linear relationship — that’s what are inferential statistics work would indicate — but there was none in the population from which the sampling was done; there would be no way for us to recognize the error except to repeat the experiment — the principal of research reproducibility — with a different sample.
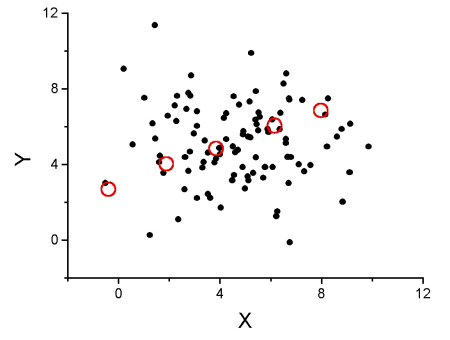
Figure 1. Scatterplot of hypothetical x,y data for which the researcher may obtain a statistically significant linear fit to sample of data from population in which null hypothesis is true relationship between x and y.
So in conclusion, you must keep in mind the meaning of statistical significance in the context of statistical inference: it is inference done on a background of random chance, the chance that sampling from the population leads to a biased sample of subjects.
Note 1. If you think about what I did with this data for Figure 1 graph, purposely selecting data that showed a linear relationship between Y and X (red circles), then you should recognize this as an example of data dredging or p-hacking, cf. discussion in Head et al (2015); Stefan and Schönbrodt (2023). However, the graph is supposed to be read as if we could do a census and therefore have full knowledge of the true relationship between y and x. The red circles indicate the chance that sampling from a population may sometimes yield incorrect conclusions.
Tests of coefficients
One criterion for a good model is that the coefficients in the model, the intercept and the slope(s) are all statistically significant.
For the statistical of the slope, b1, we generally treat the test as a two-tailed test of the null hypothesis that the regression slope is equal to zero.
H0 : b1 = 0 vs. HA : b1 ≠ 0
Similarly, for the statistical of the intercept, b0, we generally treat the test as a two-tailed test of the null hypothesis that the Y-intercept is equal to zero.
H0 : b0 = 0 vs. HA : b0 ≠ 0
For both slope and intercept we use t-statistics.
![]()
We’ll illustrate the tests of the slope and intercept by letting R and Rcmdr do the work. You’ll find this simple data set at the bottom of this page (scroll or click here). The first variable is the number of matings, the second is the size of the paired female, and the third is the size of the paired male. All body mass were in grams.
R code
After loading the worksheet into R and Rcmdr, begin by selecting
Rcmdr: Statistics → Fit model → Linear Regression
Note that more than one predictor can be entered, but only one response (dependent) variable may be selected (Fig 2).
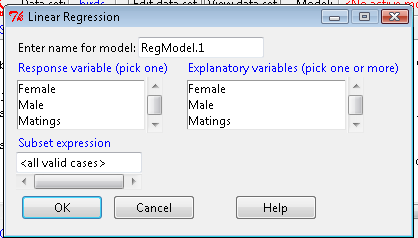
Figure 2. Screenshot linear regression menu. More than explanatory (predictor or independent) variables may be selected, but only one response (dependent) variable may be selected.
This procedure will handle simple and multiple regression problems. But before we go further, answer these two questions for the data set.
Question 1. What is the Response variable?
Question 2. What is the Explanatory variable?
Answers. See below in the R output to see if you were correct!
If there is only one predictor, then this is a Simple Linear Regression; if more than one predictor is entered, then this is a Multiple Linear Regression. We’ll get some more detail, but for now, identify and evaluate the test (p-value) of the slope coefficient (labeled after the name of the predictor variable), the test of the intercept coefficient, and some new stuff.
R output
RegModel.1 <- lm(Matings ~ Female, data=birds)
summary(RegModel.1)
Call: lm(formula = Matings ~ Female, data = birds)
Residuals:
Min 1Q Median 3Q Max
-2.32805 -1.59407 -0.04359 1.77292 2.67195
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -8.6175 3.5323 -2.440 0.02528 *
Female 0.3670 0.1042 3.524 0.00243 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.774 on 18 degrees of freedom
Multiple R-squared: 0.4082, Adjusted R-squared: 0.3753 F-statistic: 12.42 on 1 and 18 DF, p-value: 0.002427
In this example, slope = 0.367 and the intercept = -8.618. The first new term we encounter is called Multiple R-squared (R2) — it’s also called the coefficient of determination. It’s the ratio of the sum of squares due to the regression to the total sums of squares. R2 ranges from zero to 1, with a value of 1 indicating a perfect fit of the regression to the data.
Note 2. If you are looking for a link between correlation and simple linear regression, then here it is: R2 is the square of the product-moment correlation, r. Thus, r = √R2 see Chapter 17.2).
We introduced ![]() in Chapter 17.1 . Interpretation of R2 goes like this: If R2 is close to zero, then the regression model does not explain much of the variation in the dependent variable; conversely, if R2 is close to one, then the regression model explains a lot of the variation in the dependent variable.
in Chapter 17.1 . Interpretation of R2 goes like this: If R2 is close to zero, then the regression model does not explain much of the variation in the dependent variable; conversely, if R2 is close to one, then the regression model explains a lot of the variation in the dependent variable.
Did you get the Answers?
Answer 1: Number of matings
Answer 2: Size of females
Interpreting the output
Recall that
![]()
then
![]()
From the output we see that R2 was 0.408, which means that about 40% of the variation in numbers of matings may be explained by size of the females alone.
To complete the analysis get the ANOVA table for the regression.
Rcmdr: Models → Hypothesis tests → ANOVA table…
> Anova(RegModel.1, type="II")
Anova Table (Type II tests)
Sum Sq Df F value Pr(>F)
Female 39.084 1 12.415 0.002427 **
Residuals 56.666 18
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
With only one predictor (explanatory) variable, note that the regression test in the ANOVA is the same (has the same probability vs. the null hypothesis) as the test of the slope.
Test two slopes
A more general test of the null hypothesis involving the slope might be
H0 : b1 = b vs. HA : b1 ≠ b
where b can be any value, including zero. Again, t-test would then be used to conduct the test of the slope, where the t-test would have the form
![]()
where SEb1-b2 is the standard error of the difference between the two regression coefficients. We saw something similar to this value back when we did a paired t-test. To obtain sb1–b2, we need the pooled residual mean square and the squared sums of the X values for each of our sample.
First, the pooled (hence the subscript p) residual mean square is calculated as
![]()
where resSS and resDF refer to residual sums of squares and residual degrees of freedom for the first (1) and second (2) regression equations.
Second, the standard error of the difference between regression coefficients (squared!!) is calculated as
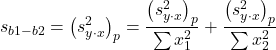
where the subscript “1” and “2” refer to the X values from the first sample (eg, the body size values for the males) and the second sample (eg, the body size values for the females).
Note 3. To obtain the squared sum in R and Rcmdr, use the Calc function (eg, two sum the squared X values for the females, use SUM(‘Female’*’Female’)).
We can then use our t-test, with the degrees of freedom now
![]()
Alternatively, the t-test of two slopes can be written as
![]()
with again ![]() .
.
In this way, we can see a way to test any two slopes for equality. This would be useful if we wanted to compare two samples (eg, males and females) and wanted to see if the regressions were the same (eg, metabolic rate covaried with body mass in the same way — that is, the slope of the relationship was the same). This situation arises frequently in biology. For example, we might want to know if male and female birds have different mean field metabolic rates, in which case we might be tempted to use a one-way ANOVA or t-test (since one factor with two levels). However, if males and females also differ for body size, then any differences we might see in metabolic rate could be due to differences in metabolic rate or to differences in the covariable body size. The test is generally referred to as the analysis of covariance (ANCOVA), which is the subject for Chapter 17.6. In brief, ANCOVA allows you to test for mean differences in traits like metabolic rate between two or more groups, but after first accounting for covariation due to another variable (eg, body size). However, ANCOVA makes the assumption that relationship between the covariable and the response variable is the same in the two groups. This is the same as saying that the regression slopes are the same. Let’s proceed to see how we can compare regression slopes, then move to a more general treatment in Chapter 17.6.
Example
For a sample of 13 tadpoles (Rana pipiens), hatched in the laboratory, oxygen consumption at rest was recorded with a YSI meter, model 51B, and a YSI oxygen – temperature probe (Yellow Spring Instruments Co., Yellow Springs, Ohio). Individual tadpoles were placed into a mason jar (volume 885 ml), with a magnetic stir bar housed in a slotted Petri dish glued to the bottom of the jar to gently stir the water. Water samples from the aquarium were used and petroleum jelly served as a sealant once the vessel had been fitted with the probe. Oxygen consumption was recorded in ppm and ![]() values (ml O2 STP · hr-1). The tadpoles were scored for developmental stage by Gosner’s (1960) criteria. The tadpoles ranged from Gosner stage 35 to 44.
values (ml O2 STP · hr-1). The tadpoles were scored for developmental stage by Gosner’s (1960) criteria. The tadpoles ranged from Gosner stage 35 to 44.
Note 4. Gosner stages 1 through 25 include the embryonic stages; By stage 31, toes develop on the limb buds; Following stage 40, substantial body changes are observed in the growing tadpole, transitioning from tailed tadpole to tail-less adult form. Complete metamorphosis is assigned Gosner stage 45.
Figure 3. Pearson Scott Foresman, Public domain, via Wikimedia Commons
The question we addressed: metamorphosis of Anuran tadpoles to adults (frogs) likely involves increased metabolic demands, demands that may exceed simple allometric predictions based on increase in body size during development. Thus, we predicted that mass-specific metabolic rate for metamorphic frogs would exceed mass-specific metabolism of pre-metamorphic frogs.
M. Dohm unpublished results from my undergraduate days (1986!), you’ll find this simple data set at the bottom of this page (scroll or click here). For simplicity, we grouped all tadpoles between stages 35 and 38 as Stage I and all tadpoles between 39 and 44 as stage II. No tadpoles who completed metamorphosis were included in the dataset. It shouldn’t surprise you — with such a small sample size, the project lacked power to test the hypothesis, but many such studies, it’s worth looking at in the case anyway because the prevailing sentiment is that the metabolic cost of metamorphosis is significant — in other words, we expect a large effect size — and remains an ongoing research effort (see Burraco et al 2019).
You should confirm your self, but the slope of the regression equation of oxygen consumption (ml O2/hour) on body mass (g) was 444.95 (with SE = 65.89). Plot of data shown in Figure 4.
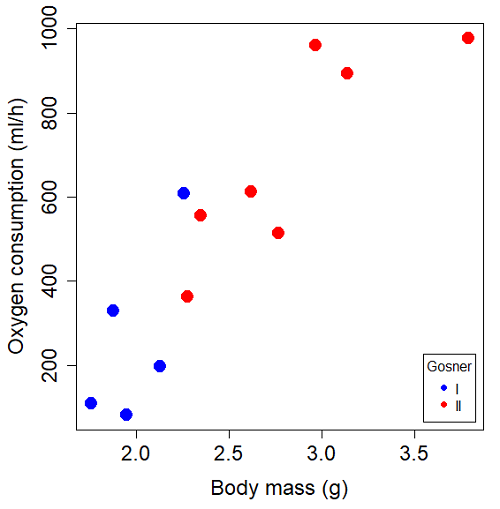
Figure 4. Scatterplot of oxygen consumption by tadpoles (blue: Gosner developmental stage I [35 – 38]; red: Gosner developmental stage II [39 – 44]), vs body mass (g).
The project looked at whether metabolism as measured by oxygen consumption was consistent across two developmental stages. Metamorphosis in frogs and other amphibians represents profound reorganization of the organism as the tadpole moves from water to air. Thus, we would predict some cost as evidenced by change in metabolism associated with later stages of development. Figure 5 shows a box plot of tadpole oxygen consumption by Gosner (1960) developmental stage.
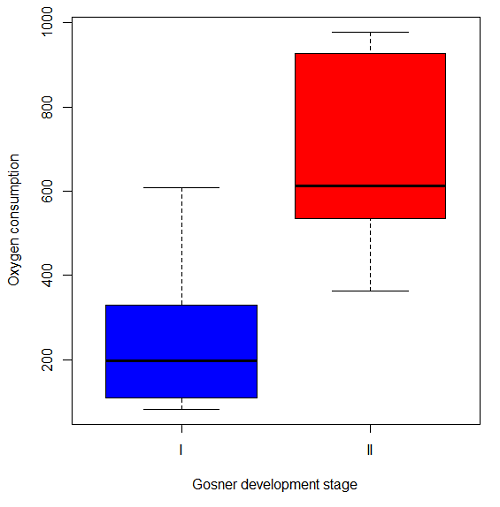
Figure 5. Boxplot of oxygen consumption by Gosner developmental stages (blue: stage I; red: stage 2).
Looking at Figure 4 we see a trend consistent with our prediction; developmental stage may be associated with increased metabolism. However, older tadpoles also tend to be larger, and the plot in Figure 4 does not account for that. Thus, body mass is a confounding variable in this example. There are several options for analysis here (eg, ANCOVA), but one way to view this is to compare the slopes for the two developmental stages. While this test does not compare the means, it does ask a related question: is there evidence of change in rate of oxygen consumption relative to body size between the two developmental stages? The assumption that the slopes are equal is a necessary step for conducting the ANCOVA, which we describe in Chapter 17.6.
So, divide the data set into two groups by developmental stage (12 tadpoles could be assigned to one of two developmental stages; one tadpole was at a lower Gosner stage than the others and so is dropped from the subset).
Gosner stage I (stages 35 – 39)
| Body mass | |
| 1.76 | 109.41 |
| 1.88 | 329.06 |
| 1.95 | 82.35 |
| 2.13 | 198 |
| 2.26 | 607.7 |
Gosner stage II (stage 40 – 44)
| Body mass | |
| 2.28 | 362.71 |
| 2.35 | 556.6 |
| 2.62 | 612.93 |
| 2.77 | 514.02 |
| 2.97 | 961.01 |
| 3.14 | 892.41 |
| 3.79 | 976.97 |
The slopes and standard errors were
| Gosner Stage I | Gosner stage II | |
| slope | 750.0 | 399.9 |
| standard error of slope | 444.6 | 111.2 |
Are the two slopes equal?
Looking at the table, we would say, no. slopes look different (750 vs 399.9). However, the errors are large and, given this is a small data set, we need to test statistically; are the slopes indistinguishable (Ho: bI = bII), where bI is the slope for the Gosner Stage I subset and bII is the slope for the Gosner Stage II subset?
To use our tests discussed above, we need sum of squared X values for Gosner Stage I and sum of squared for Gosner stage II results. We can get these from the ANOVA tables. Recall that after applying
ANOVA regression Gosner Stage I, R work
Anova(gosI.1, type="II")
Anova Table (Type II tests)
Response: VO2
Sum Sq Df F value Pr(>F)
Body.mass 89385 1 2.8461 0.1902
Residuals 94220 3
ANOVA regression Gosner Stage II
Anova(gosII.1, type="II")
Anova Table (Type II tests)
Response: VO2
Sum Sq Df F value Pr(>F)
Body.mass 258398 1 12.935 0.0156 *
Residuals 99882 5
Rcmdr: Models – Compare model coefficients..
compareCoefs(gosI.1, gosII.1)
Calls:
1: lm(formula = VO2 ~ Body.mass, data = gosI)
2: lm(formula = VO2 ~ Body.mass, data = gosII)
Model 1 Model 2
(Intercept) -1232 -441
SE 891 321
Body.mass 750 400
SE 445 111
SSgosnerI = 89385
SSgosnerII = 258398
We also need the residual Mean Squares (SSresid/DF resid) from the ANOVA tables
MSgosnerI = 94220/3 = 31406.67
MSgosnerII = 99882/5 = 19976.4
[under construction] Therefore, the pooled residual MS (s2x.y)p = nnn and the pooled SE of the difference sb1-b2 = nnn using the formulas above.
Now, we plug in the values I can get a t-test = (750 – 444.6)/nnn = nnn
The DF for this t-test are n1 + n2 – 4 = 4 + 6 – 4 = 6.
Using Table of Student’s t distribution (Appendix), I find the two-tailed critical value for t at alpha = 5% with DF = 6 is equal to 3.758. Since nn >> 3.758, we conclude that the two slopes are statistically different and P < 0.001 that the null hypothesis (Ho: bw = ball) was true. (edit: fix nn).
Questions
1. Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
2. Metabolic rates like oxygen consumption over time are well-known examples of allometric relationships. That is, many biological variables (eg, ![]() is related as a · Mb, where M is body mass, b is scaling exponent (the slope!), and a is a constant (the intercept!)), and best evaluated on log-log scale. Redo the linear regression of oxygen consumption vs. body mass for the tadpoles, but this time, apply log10-transform to
is related as a · Mb, where M is body mass, b is scaling exponent (the slope!), and a is a constant (the intercept!)), and best evaluated on log-log scale. Redo the linear regression of oxygen consumption vs. body mass for the tadpoles, but this time, apply log10-transform to VO2 and to Body.mass. For example
Data in this page, bird matings
| Body.Mass | Matings |
|---|---|
| 29 | 0 |
| 29 | 2 |
| 29 | 4 |
| 32 | 4 |
| 32 | 2 |
| 35 | 6 |
| 36 | 3 |
| 38 | 3 |
| 38 | 5 |
| 38 | 8 |
| 40 | 6 |
Data in this page, Oxygen consumption, ![]() , of Anuran tadpoles
, of Anuran tadpoles
| Gosner | Body mass | VO2 |
|---|---|---|
| NA | 1.46 | 170.91 |
| I | 1.76 | 109.41 |
| I | 1.88 | 329.06 |
| I | 1.95 | 82.35 |
| I | 2.13 | 198 |
| II | 2.28 | 362.71 |
| I | 2.26 | 607.7 |
| II | 2.35 | 556.6 |
| II | 2.62 | 612.93 |
| II | 2.77 | 514.02 |
| II | 2.97 | 961.01 |
| II | 3.14 | 892.41 |
Gosner refers to Gosner (1960), who developed a criteria for judging metamorphosis staging.
Quiz Chapter 17.5
Testing regression coefficients
Chapter 17 contents
- Introduction
- Simple Linear Regression
- Relationship between the slope and the correlation
- Estimation of linear regression coefficients
- OLS, RMA, and smoothing functions
- Testing regression coefficients
- ANCOVA – Analysis of covariance
- Regression model fit
- Assumptions and model diagnostics for Simple Linear Regression
- References and suggested readings (Ch17 & 18)
17.4 – OLS, RMA, and smoothing functions
Introduction
OLS or ordinary least squares is the most commonly used estimation procedure for fitting a line to the data. For both simple and multiple regression, OLS works by minimizing the sum of the squared residuals. OLS is appropriate when the linear regression assumptions LINE apply. In addition, further restrictions apply to OLS including that the predictor variables are fixed and without error. OLS is appropriate when the goal of the analysis is to retrieve a predictive model. OLS describes an asymmetric association between the predictor and the response variable: the slope bX for Y ~ bXX will generally not be the same as the slope bY for X ~ bYY.
OLS is appropriate for assessing functional relationships (i.e., inference about the coefficients) as long as the assumptions hold. In some literature, OLS is referred to as a Model I regression.
Generalized Least Squares
Generalized linear regression is an estimation procedure related to OLS but can be used either when variances are unequal or multicollinearity is present among the error terms.
Weighted Least Squares
A conceptually straightforward extension of OLS can be made to account for situation where the variances in the error terms are not equal. If the variance of Yi varies for each Xi, then a weighting function based on the reciprocal of the estimated variance may be used.
![]()
then, instead of minimizing the squared residuals as in OLS, the regression equation estimates in weighted least squares minimizes the squared residuals summed over the weights.
![]()
Weighted least squares is a form of generalized least squares. In order to estimate wi, however, multiple values of Y for each observed X must be available.
Reduced Major Axis
There are many alternative methods available when OLS may not be justified. These approaches, collectively, may be called Model II regression methods. These methods are invariably invoked in situations in which both Y and X variables have random error associated with them. In other words, the OLS assumption that the predictor variables are measured without error is violated. Among the more common methods is one called Reduced Major Axis or RMA.
Smoothing functions
data set: atmospheric carbon dioxide (CO2) readings Mauna Loa. Source: https://gml.noaa.gov/ccgg/trends/data.html
Fit curves without applying a known formula. This technique is called smoothing and, while there are several versions, the technique involves taking information from groups of observations — weighted averaged — and using these groups to estimate how the response variable changes with values of the independent variable. Smoothing is used to help reveal patterns, to emphasize trends by reducing noise — clearly, caution needs to be employed as smoothing necessarily hides outlier data, which can themselves be important. Smoothing techniques by name include kernel, loess, and spline. Default in the scatter plot command is loess.
CO2 in parts per million (ppm) plotted by year from 1958 to 2014 the first CO2 readings were recorded in April 1958 (Fig 1).
Note 1: When I worked on this set, the last data available for this plot was April 2014. CO2 421.86 ppm December 2023, 399.08 ppm December 2014 — a 5.7% increase. https://www.esrl.noaa.gov/gmd/ccgg/trends/
A few words of explanation for Figure 1. The green line shows the OLS line, the red line shows the loess smoothing with a smoothing parameter of 0.5 (in Rcmdr the slider value reads “5”).
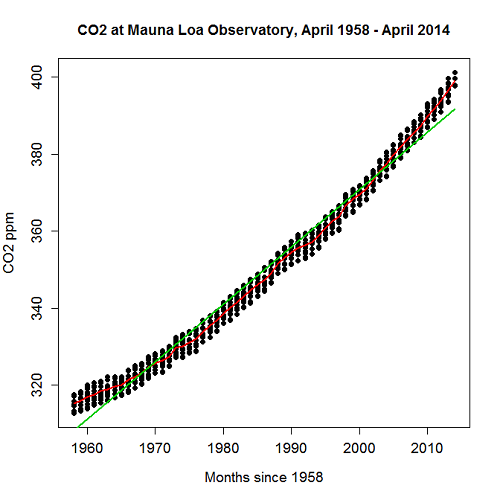
Figure 1. CO2 in parts per million (ppm) plotted by year from 1958 to 2014
R command was started with option settings available in Rcmdr context menu for scatterplot, then additional commands were added
scatterplot(CO2~Year, reg.line=lm, grid=FALSE, smooth=TRUE, spread=FALSE, boxplots=FALSE, span=0.05, lwd=2, xlab="Months since 1958", ylab="CO2 ppm", main="CO2 at Mauna Loa Observatory, April 1958 - April 2014", cex=1, cex.axis=1.2, cex.lab=1.2, pch=c(16), data=StkCO2MLO)
The next plot is for ppm CO2 by month for the year 2013. The plot shows the annual cycle of atmospheric CO2 in the northern hemisphere.
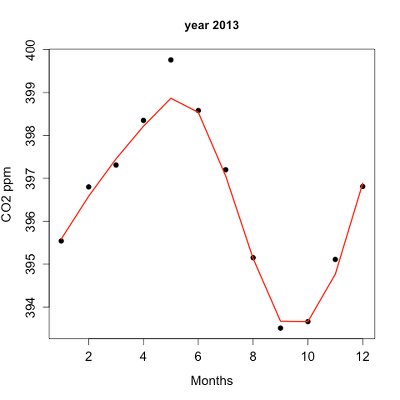
Figure 2. Plot of ppm CO2 by month for the year 2013.
Again, the smoothing parameter was set to 0.5 and the loess function is plotted in red (Fig 2).
Loess is an acronym short for local regression. Loess is a weighted least squares approach which is used to fit linear or quadratic functions of the predictors at the centers of neighborhoods. The radius of each neighborhood is chosen so that the neighborhood contains a specified percentage of the data points. This percentage of data points is referred to as the smoothing parameter and this parameter may differ for different neighborhoods of points. The idea of loess, in fact, any smoothing algorithm, is to reveal pattern within a noisy sequence of observations. The smoothing parameter can be set to different values, between 0 and 1 is typical.
Note 2: Noisy data in this context refers to data comes with random error independent of the true signal, i.e., noisy data has low signal-to-noise ratio. The concept is most familiar in communication.
To get a sense of what the parameter does, Figure 3 takes the same data as in Figure 2, but with different values of the smoothing parameter (Fig 3).
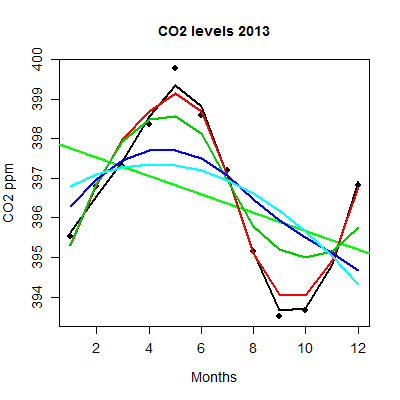
Figure 3. Plot with different smoothing values (0.5 to 10.0).
key:
| parameter | color |
| 0.5 | black |
| 0.75 | red |
| 1.0 | dark green |
| 2.0 | blue |
| 10.0 | light blue |
The R code used to generate Figure 3 plot was
spanList = c(0.5, 0.75, 1, 2, 10)
reg1 = lm(ppm~Month)
png(filename = "RplotCO2mo.png", width = 400, height = 400, units = "px", pointsize = 12, bg = "white")
plot(Month,ppm, cex=1.2, cex.axis=1.2, cex.lab=1.2, pch=c(16), xlab="Months", ylab="CO2 ppm", main="CO2 levels 2013")
abline(reg1,lwd=2,col="green")
for (i in 1:length(spanList))
{
ppm.loess <- loess(ppm~Month, span=spanList[i], Dataset)
ppm.predict <- predict(ppm.loess, Month)
lines(Month,ppm.predict,lwd=2,col=i)
}
Note 3: This is our first introduction to use of a “for” loop.
The CO2 data constitutes a time series. Instead of loess, a simple moving average would be a more natural way to reveal trends. In principle, take a set of nearby points (odd number of points best, keeps the calculation symmetric) and calculate the average. Next, shift the points by a specified time interval (eg, 7 days), and recalculate the average for the new set of points. See Chapter 20.5 for Time series analysis.
Questions
- Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
- This is bio-statistics, so I gotta ask: What ecological/environmental process explains the shape of the relationship between ppm CO2 and months of the year as shown in Figure 2? Hint: NOAA Global Monitoring Laboratory responsible for the CO2 data is located at Mauna Loa Observatory, Hawaiʻi (lat: 19.52291, lon: -155.61586).
- As I wrote this question (January 2022), we were 22 months since WHO declared Covid-19 a pandemic (CDC timeline). Omicron variant is now dominant; Daily case counts State of Hawaiʻi from 1 November 2021 to 15 January 2022 reported in data set table.
- Make plot like Figure 2 (days instead of months)
- Apply different loess smoothing parameters and re-plot the data. Observe and describe the change to the trend between case reports and days.
Quiz Chapter 17.4
OLS, RMA, and smoothing functions
Data set
Covid-19 cases reported State of Hawaiʻi from 1 November 2021 to 15 January 2022 (data extracted from Wikipedia)
| Date | Cases reported |
|---|---|
| 11/01/21 | 69 |
| 11/02/21 | 38 |
| 11/03/21 | 176 |
| 11/04/21 | 112 |
| 11/05/21 | 124 |
| 11/06/21 | 97 |
| 11/07/21 | 134 |
| 11/08/21 | 94 |
| 11/09/21 | 79 |
| 11/10/21 | 142 |
| 11/11/21 | 130 |
| 11/12/21 | 138 |
| 11/13/21 | 81 |
| 11/14/21 | 0 |
| 11/15/21 | 146 |
| 11/16/21 | 93 |
| 11/17/21 | 142 |
| 11/18/21 | 226 |
| 11/19/21 | 206 |
| 11/20/21 | 218 |
| 11/21/21 | 107 |
| 11/22/21 | 92 |
| 11/23/21 | 52 |
| 11/24/21 | 115 |
| 11/25/21 | 77 |
| 11/26/21 | 27 |
| 11/27/21 | 135 |
| 11/28/21 | 169 |
| 11/29/21 | 71 |
| 11/30/21 | 79 |
| 12/01/21 | 108 |
| 12/02/21 | 126 |
| 12/03/21 | 125 |
| 12/04/21 | 124 |
| 12/05/21 | 148 |
| 12/06/21 | 90 |
| 12/07/21 | 55 |
| 12/08/21 | 72 |
| 12/09/21 | 143 |
| 12/10/21 | 170 |
| 12/11/21 | 189 |
| 12/12/21 | 215 |
| 12/13/21 | 150 |
| 12/14/21 | 214 |
| 12/15/21 | 282 |
| 12/16/21 | 395 |
| 12/17/21 | 797 |
| 12/18/21 | 707 |
| 12/19/21 | 972 |
| 12/20/21 | 840 |
| 12/21/21 | 707 |
| 12/22/21 | 961 |
| 12/23/21 | 1511 |
| 12/24/21 | 1828 |
| 12/25/21 | 1591 |
| 12/26/21 | 2205 |
| 12/27/21 | 1384 |
| 12/28/21 | 824 |
| 12/29/21 | 1561 |
| 12/30/21 | 3484 |
| 12/31/21 | 3290 |
| 01/01/22 | 2710 |
| 01/02/22 | 3178 |
| 01/03/22 | 3044 |
| 01/04/22 | 1592 |
| 01/05/22 | 2611 |
| 01/06/22 | 4789 |
| 01/07/22 | 3586 |
| 01/08/22 | 4204 |
| 01/09/22 | 4578 |
| 01/10/22 | 3875 |
| 01/11/22 | 2929 |
| 01/12/22 | 3512 |
| 01/13/22 | 3392 |
| 01/14/22 | 3099 |
| 01/15/22 | 5977 |
Chapter 17 contents
- Introduction
- Simple Linear Regression
- Relationship between the slope and the correlation
- Estimation of linear regression coefficients
- OLS, RMA, and smoothing functions
- Testing regression coefficients
- ANCOVA – analysis of covariance
- Regression model fit
- Assumptions and model diagnostics for Simple Linear Regression
- References and suggested readings (Ch17 & 18)
17.3 – Estimation of linear regression coefficients
Introduction
In discussing correlations we made the distinction between inference, testing the statistical significance of the estimate, and the process of getting the estimate of the parameter itself. Estimating parameters is possible for any data set; whether or not the particular model is a good and useful model is another matter. Statisticians speak about the fit of a model… that a model with one or more independent predictor variables explains a substantial amount of the variation in the dependent variable, that it describes the relationship between the predictors and the dependent variable without bias.
Note 1. We introduced the concept of statistical fit in our discussion of Chi-square, goodness of fit, Chapter 9.1.
A number of tools have been developed to assess model fit. For starters, I’ll list just two ways you can approach whether or not a linear model fits your data ore requires some intervention on your part.
Assess fit of a linear model
Recall our R output from the regression of Number of Matings on Body Mass from the bird data set. We used the linear model function.
LinearModel.1 <- lm(Matings ~ Body.Mass, data=bird_matings)
summary(LinearModel.1)
Call: lm(formula = Matings ~ Body.Mass, data = bird_matings)
Residuals: Min 1Q Median 3Q Max
-2.29237 -1.34322 .-0.03178..1.33792 .2.70763
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -8.4746 4.6641 -1.817 0.1026
Body.Mass 0.3623 0.1355 2.673 0.0255 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.776 on 9 degrees of freedom
Multiple R-squared: 0.4425, Adjusted R-squared: 0.3806
F-statistic: 7.144 on 1 and 9 DF, p-value: 0.02551
Request R to print the ANOVA table.
Anova(RegModel.1, type="II")
Anova Table (Type II tests)
Response: Matings
Sum Sq Df F value Pr(>F)
Body.Mass 22.528 1 7.1438 0.02551 *
Residuals 28.381 9
With a little rounding we have the following statistical model
![]()
and in English, Number of matings equals Body.mass multiplied by 0.36 then subtract 8.5; the intercept was -8.5, the slope was 0.36.
Note 2. The results of the regression analyses were stored in the object called “LinearModel.1“. This is a nice feature of Rcmdr — it automatically provides an object name for you. Note that with each successive run of the linear model function via Rcmdr that it will change the object name by adding numbers successively. For example, after LinearModel.1 the next run of lm() in Rcmdr will automatically be called “LinearModel.2” and so on. In your own work you may specify the names of the objects directly or allow Rcmdr to do it for you, but do keep track of the object names!
From the R output we see that the estimate of the slope was +0.36, statistically different from zero (p = 0.025). The intercept was -8.5, but not statistically significant (p = 0.103), which means the intercept may be zero.
As a general rule, if you make an estimate of a parameter or coefficient, then you should provide a confidence interval of the form.
estimate + critical value X standard error of the estimate
Note 3. Reminder: Approximate 95% CI can be obtained by + twice the standard error for the coefficient.
For confidence interval of regression slope we have a couple of options in R
Option 1.
confint(LinearModel.1, 'Body.Mass', level=0.95) 2.5 % 97.5 % Body.Mass 0.05565899 0.6689173
Option 2.
Goal: Extract the coefficients from the output of the linear model and calculate the approximate SE with nine degrees of freedom. This is the big advantage of saving output from functions as objects. Typically, much more information is about the results are available, and, additionally, can be retrieved for additional use. Extracting coefficients from the objects is the best option, but does come with a learning curve. Let’s get started.
First, what information is available in the linear model output beyond the default information? To find out, use the names() function
names(LinearModel.1)
R output
[1] "coefficients" "residuals" "effects" "rank" [5] "fitted.values" "assign" "qr" "df.residual" [9] "xlevels" "call" "terms" "model"
Another way is to use the summary() function call.
summary(LinearModel.1)$coefficients Estimate Std. Error t value Pr(>|t|) (Intercept) -8.4745763 4.6640856 -1.816986 0.10259256 Body.Mass 0.3622881 0.1355472 2.672781 0.02550595
How can we get just the standard error for the slope? Note that the estimates are reported in a 2X4 matrix like so
| 1,1 | 1,2 | 1,3 | 1,4 |
| 2,1 | 2,2 | 2,3 | 2,4 |
Therefore, to get the standard error for the slope we identify that it is stored in cell 2,2 of the matrix and we write
summary(LinearModel.1)$coefficients[2,2]
which returns
[1] 0.1355472
Let’s use this information to calculate confidence intervals
slp=summary(LinearModel.1)$coefficients[2,1] slpErrs=summary(LinearModel.1)$coefficients[2,2] slp + c(-1,1)*slpErrs*qt(0.975, 9)
where qt() is the quantile function for the t distribution and “9” is the degrees of freedom from the regression. Results follow
coef + c(-1,1)*errs*qt(0.975, 9) [1] 0.05565899 0.66891728
And for the intercept
int=summary(LinearModel.1)$coefficients[1,1] intErrs=summary(LinearModel.1)$coefficients[1,2] int + c(-1,1)*intErrs*qt(0.975, 9)
Results
int + c(-1,1)*intErrs*qt(0.975, 9) [1] -19.025471 2.076318
In conclusion, part of fitting a model includes reporting the estimates of the coefficients (model parameters). And, in general, when estimation is performed, reporting of suitable confidence intervals are expected.
Extract additional statistics from R’s linear model function
The summary() function is used to report the general results from ANOVA and linear model function output in R software, but additional functions can be used to extract the rest of the output, e.g., coefficient of determination. To complete our example of extracting information from the summary() function, we next turn to summary.lm() function to see what is available.
At the R prompt type and submit
summary.lm(LinearModel.1)
returns the following R output
Call: lm(formula = Matings ~ Body.Mass, data = bird_matings) Residuals: Min 1Q Median 3Q Max -2.29237 -1.34322 -0.03178 1.33792 2.70763 Coefficients: Estimate Std. Error t value Pr(>|t|) (Intercept) -8.4746 4.6641 -1.817 0.1026 Body.Mass 0.3623 0.1355 2.673 0.0255 * --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 Residual standard error: 1.776 on 9 degrees of freedom Multiple R-squared: 0.4425, Adjusted R-squared: 0.3806 F-statistic: 7.144 on 1 and 9 DF, p-value: 0.02551
Looks exactly like the output from summary(). Let’s look at what is available in the summary.lm() function
names(summary.lm(LinearModel.1)) [1] "call" "terms" "residuals" "coefficients" [5] "aliased" "sigma" "df" "r.squared" [9] "adj.r.squared" "fstatistic" "cov.unscaled"
We see some information we got from summary(), e.g., “coefficients”. If we interrogate the name coefficients like so
summary.lm(LinearModel.1)$coefficients
we get
summary.lm(LinearModel.1)$coefficients Estimate Std. Error t value Pr(>|t|) (Intercept) -8.4745763 4.6640856 -1.816986 0.10259256 Body.Mass 0.3622881 0.1355472 2.672781 0.02550595
which, again, is a 2X4 matrix (see above)
So to get the standard error for the slope we identify that it is stored in cell 2,2 of the matrix and call it LinearModel.1$coefficients[2,2].
Questions
1. Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
2. What does a small standard error of the slope for a SLR model tell us?
3. For slope, intercept, for just about any statistical estimate, why are we obligated to provide a confidence interval?
Quiz Chapter 17.3
Estimation of linear regression coefficients
Chapter 17 contents
- Introduction
- Simple Linear Regression
- Relationship between the slope and the correlation
- Estimation of linear regression coefficients
- OLS, RMA, and smoothing functions
- Testing regression coefficients
- ANCOVA – analysis of covariance
- Regression model fit
- Assumptions and model diagnostics for Simple Linear Regression
- References and suggested readings (Ch17 & 18)
17.2 – Relationship between the slope and the correlation
Introduction
Product moment correlation is used to indicate the direction and the strength of the linear association between two ratio-scale variables; the slope tells you the rate of change between the two variables. When the correlation is negative, the slope will be negative; when correlation is positive, so too will the slope.
As you might suspect, there is a mathematical relationship between the product moment correlation, r, and the regression slope, b1. We haven’t spent much time explaining the equations presented in this text, but correlation and linear regression are such important tools it’s worth a closer look.
Recall from our discussion in Chapter 16.1, the equation of the correlation is
![]()
where the numerator is termed the covariance between X and Y and the denominator contains the standard deviations of X and Y variables. We can say the at the covariance is standardized by the variability in X and Y. In contrast, the regression slope is equal to the covariance divided by the variance in X.
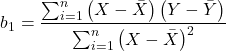
Thus, with a little algebra, we can see that the slope and correlation are equal to each other as
![]()
This should drive home the following statistical reasoning point. You can always calculate a slope from a correlation, but recall that correlation analysis is intended as a test of the hypothesis of a linear association between variables for which cause and effect model — though perhaps reasonable — should be implied. Just because it is mathematically possible does not mean the analysis is correct for the problem.
Coefficient of determination and the Product moment correlation
![]() , the coefficient of determination, was introduced in the last chapter. It’s the ratio of variation of the data explained by the linear regression model divided by the total variation in the data. Values range from 0% to 100% — it’s a measure of fit, how well the data are described by a line. Note that part of our description for
, the coefficient of determination, was introduced in the last chapter. It’s the ratio of variation of the data explained by the linear regression model divided by the total variation in the data. Values range from 0% to 100% — it’s a measure of fit, how well the data are described by a line. Note that part of our description for ![]() , the product moment correlation — strength of the linear association — is simply another way to describe model fit. Thus,
, the product moment correlation — strength of the linear association — is simply another way to describe model fit. Thus,
![]()
Of course, we lose the direction information by squaring the correlation.
Questions
- Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
- If the correlation is 0.6,
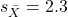 , and
, and 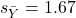 , what is the slope estimate?
, what is the slope estimate?
Quiz Chapter 17.2
Relationship between the slope and the correlation
Chapter 17 contents
- Introduction
- Simple Linear Regression
- Relationship between the slope and the correlation
- Estimation of linear regression coefficients
- OLS, RMA, and smoothing functions
- Testing regression coefficients
- ANCOVA – analysis of covariance
- Regression model fit
- Assumptions and model diagnostics for Simple Linear Regression
- References and suggested readings (Ch17 & 18)
17.1 – Ordinary least squares regression
Introduction
Linear regression is a toolkit for developing linear models of cause and effect between a ratio scale data type, response or dependent variable, often labeled “Y,” and one or more ratio scale data type, predictor or independent variables, X. Like ANOVA, linear regression is a special case of the general linear model. Regression and correlation both test linear hypotheses: we state that the relationship between two variables is linear (the alternative hypothesis) or it is not (the null hypothesis). The difference?
- Correlation is a test of linear association (are variables correlated, we ask?), but are not tests of causation: we do not imply that one variable causes another to vary, even if the correlation between the two variables is large and positive, for example. Correlations are used in statistics on data sets not collected from explicit experimental designs incorporated to test specific hypotheses of cause and effect.
- Linear regression is to cause and effect as correlation is to association. With regression and ANOVA, which again, are special cases of the general linear model (LM), we are indeed making a case for a particular understanding of the cause of variation in a response variable: modeling cause and effect is the goal.
We start our LM model as ![]()
where “~”, tilda, is an operator used by R in formulas to define the relationship between the response variable and the predictor variable(s).
From R Commander we call the linear model function by Statistics → Fit models → Linear model … , which brings up a menu with several options (Fig 1).
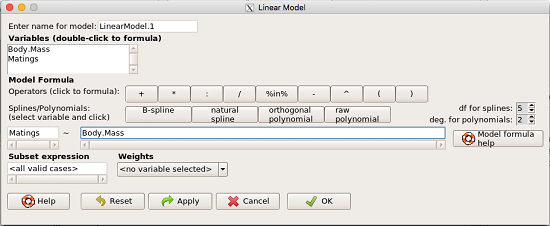
Figure 1. R commander menu interface for linear model.
Our model was
Matings ~ Body.Mass
R commander will keep track of the models created and enter a name for the object. You can, and probably should, change the object name yourself. The example shown in Figure 1 is a simple linear regression, with Body.Mass as the Y variable and Matings the X variable. No other information need be entered and one would simply click OK to begin the analysis.
Example
The purpose of regression is similar to ANOVA. We want a model, a statistical representation to explain the data sample. The model is used to show what causes variation in a response (dependent) variable using one or more predictors (independent variables). In life history theory, mating success is an important trait or characteristic that varies among individuals in a population. For example we may be interested in determining the effect of age (X1) and body size (X2) on mating success for a bird species. We could handle the analysis with ANOVA, but we would lose some information. In a clinical trial, we may predict that increasing Age (X1) and BMI (X2) causes increase blood pressure (Y).
Our causal model looks like
![]()
Let’s review the case for ANOVA first.
The response (dependent variable), the number of successful matings for each individual male bird, would be a quantitative (interval scale) variable. (Reminder: You should be able to tell me what kind of analysis you would be doing if the dependent variable was categorical!) If we use ANOVA, then factors have levels. For example, we could have several adult birds differing in age (factor 1) and of different body sizes. Age and body size are quantitative traits, so, in order to use our standard ANOVA, we would have to assign individuals to a few levels. We could group individuals by age (eg, < 6 months, 6 – 10 months, > 12 months) and for body size (eg, small, medium, large). For the second example, we might group the subjects into age classes (20-30, 30-40, etc), and by AMA recommended BMI levels (underweight < 18.5, normal weight 18.5 – 24.9, overweight 25-29.9, obese > 30).
We have not done anything wrong by doing so, but if you are a bit uneasy by this, then your intuition will be rewarded later when we point out that in most cases you are best to leave it as a quantitative trait. We proceed with the test of ANOVA, but we are aware that we’ve lost some information — continuous variables (age, body size, BMI) were converted to categories — and so we suspect (correctly) that we’ve lost some power to reject the null hypothesis. By the way, when you have a “factor” that is a continuous variable, we call it a “covariate.” Factor typically refers to a categorical explanatory (independent) variable.
We might be tempted to use correlation — at least to test if there’s a relationship between Body Mass and Number of Matings. Correlation analysis is used to measure the intensity of association between a pair of variables. Correlation is also used to to test whether the association is greater than that expected by chance alone. We do not express one as causing variation in the other variable, but instead, we ask if the two variables are related (covary). We’ve already talked about some properties of correlation: it ranges from -1 to +1 and the null hypothesis is that the true association between two variables is equal to zero. We will formalize the correlation next time to complete our discussion of the linear relationship between two variables.
But regression is appropriate here because we are indeed making a causal claim: we selected Age and Body Size, and we selected Age and BMI in the second example wish to develop a model so we can predict and maybe even advise.
Least squares regression explained
Regression is part of the general linear model family of tests. If there is one linear predictor variable, then that is a simple linear regression (SLR), also called ordinary least squares (OLS), if there are two or more linear predictor variables, then that is a multiple linear regression (MLR, Chapter 18).
First, consider one predictor variable. We begin by looking at how we might summarize the data by fitting a line to the data; we see that there’s a relationship between mass and mating success in both young and old females (and maybe in older males).
The data set was
Table 1. Our data set of number of matings by male bird by body mass (g).
| Body.Mass | Matings |
|---|---|
| 29 | 0 |
| 29 | 2 |
| 29 | 4 |
| 32 | 4 |
| 32 | 2 |
| 35 | 6 |
| 36 | 3 |
| 38 | 3 |
| 38 | 5 |
| 38 | 8 |
| 40 | 6 |
Note 1: Unfortunately, I’ve lost track where Table 1 data set came from! It’s likely a simulated set inspired by my readings back in 2005.
And a scatterplot of the data (Fig 2)
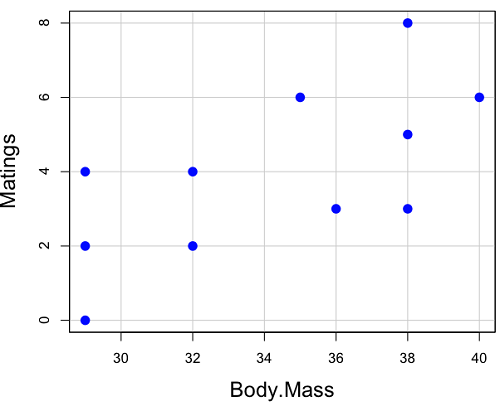
Figure 2. Number of matings by body mass (g) of the male bird.
There’s some scatter, but our eyes tell us that as body size increases, the number of matings also increases. We can go so far as to say that we can predict (imperfectly) that larger birds will have more matings. We fit the best fit line to the data and added the line to our scatterplot (Fig 3). The best fit line meets the requirements that the error about the line is minimized (see below). Thus, we would predict about six matings for a 40 g bird, but only two matings for a 28 g bird. And this is a good feature of regression, prediction, as long as used with some caution.
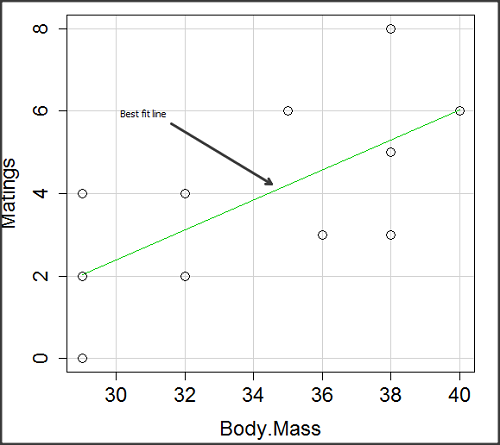
Figure 3. Same data as in Fig 2, but with the “best fit” line.
The prediction works best for the range of data for which the regression model was built. Outside the range of values, we predict with caution.
The simplest linear relationship between two variables is the SLR. This would be the parameter version (population, not samples), where
![]()
![]() = the Y-intercept coefficient and it is defined as
= the Y-intercept coefficient and it is defined as
![]()
solve for intercept by setting X = 0.
![]() = the regression coefficient (slope)
= the regression coefficient (slope)
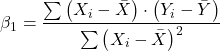
Note 2: The denominator is just our corrected sums of squares that we’ve seen many times before. The numerator is the cross-product and is referred to as the covariance.
ε = the error or “residual”
![]()
The residual is an important concept in regression. We briefly discussed “what’s left over,” in ANOVA, where an observation Yi is equal to the population mean plus the factor effect of level i plus the remainder or “error”.
In regression, we speak of residuals as the departure (difference) of an actual Yi (observation) from the predicted Y (![]() , say “why-hat”).
, say “why-hat”).
the linear regression predicts Y
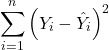
and what remains unexplained by the regression equation is called the residual
![]()
There will be as many residuals as there were observations.
But why THIS particular line? We could draw lines anywhere through the points. Well, this line is termed the “best fit“ because it is the only line that minimizes the sum of the squared deviations for all values of Y (the observations) and the predicted ![]() . The best fit line minimizes the sum of the squared residuals.
. The best fit line minimizes the sum of the squared residuals.
![]()
Note 3: More technically we justify the OLS as best fit by saying” Following the Gauss–Markov theorem, OLS estimators are the best linear unbiased estimators (BLUE). Gauss (1821) proved that the least squares method produces unbiased estimates with the smallest variance, a key result for regression analysis, while Markov (1900) rediscovered Gauss’ work and showed the conclusions held under less restrictive conditions.
Thus, like ANOVA, we can account for the total sums of squares (SSTot) as equal to the sums of squares (variation), explained by the regression model, SSreg, plus what’s not explained, what’s left over, the residual sums of squares, SSres, aka SSerror.
![]()
How well does our model explain — fit — our data? We cover this topic in Chapter 17.3 — Estimation of linear regression coefficients, but for now, we introduce ![]() , the coefficient of determination.
, the coefficient of determination.
![]()
![]() ranges from zero (0%), the linear model fails completely to explain the data, to one (100%), the linear model explains perfectly every datum.
ranges from zero (0%), the linear model fails completely to explain the data, to one (100%), the linear model explains perfectly every datum.
Models used to predict new values
Once a line has been estimated, one use is to predict new observations not previously measured!
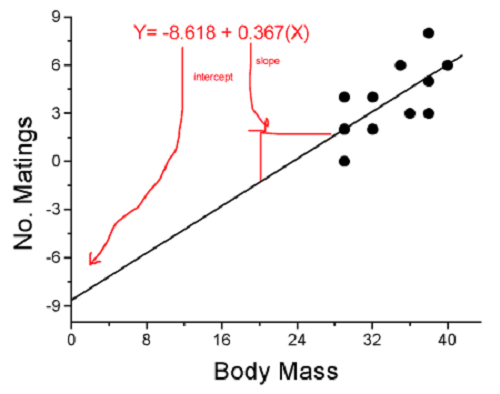
Figure 4. Figure 3 redrawn to extend the line to the Y intercept.
This is an important use of models in statistics: use an equation to fit to some data, then predict Y values from new values of X. To use the equation, simply insert new values of X into the equation, because the slope and intercept are already “known.” Then for any Xi we can determine ![]() (predicted Y value that is on the best fit regression line).
(predicted Y value that is on the best fit regression line).
This is what people do when they say
“if you are a certain weight (or BMI) you have this increased risk of heart disease”
“if you have this number of black rats in the forest you will have this many nestlings survive to leave the nest”
“if you have this much run-off pollution into the ocean you have this many corals dying”
“if you add this much enzyme to the solution you will have this much resulting product”.
R Code
We can use the drop down menu in Rcmdr to do the bulk of the work, supplemented with a little R code entered and run from the script window. Scrape data from Table 1 and save to R as bird.matings.
LinearModel.3 <- lm(Matings ~ Body.Mass, data=bird.matings)
summary(LinearModel.3)
Output from R — we’ll pull apart the important parts, color-highlights used to help orient the reader as we go.
Call: lm(formula = Matings ~ Body.Mass, data = bird.matings) Residuals: Min 1Q Median 3Q Max -2.29237 -1.34322 -0.03178 1.33792 2.70763 Coefficients: Estimate Std. Error t value Pr(>|t|) (Intercept) -8.4746 4.6641 -1.817 0.1026 Body.Mass 0.3623 0.1355 2.673 0.0255 * --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 Residual standard error: 1.776 on 9 degrees of freedom Multiple R-squared: 0.4425, Adjusted R-squared: 0.3806 F-statistic: 7.144 on 1 and 9 DF, p-value: 0.02551
What are the values for the coefficients? See Estimate .
Table 2. Coefficient estimates.
| Value | |
| -8.4746 | |
| 0.3623 |
Get the sum of squares from the ANOVA table
myAOV.full <- anova(LinearModel.3); myAOV.full
Output from R, the ANOVA table
Analysis of Variance Table Response: Matings Df Sum Sq Mean Sq F value Pr(>F) Body.Mass 1 22.528 22.5277 7.1438 0.02551 * Residuals 9 28.381 3.1535 --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Looking for ![]() , the coefficient of determination? From the R output identify Multiple R-squared. We find 0.4425. Grab the SSreg and SSTot from the ANOVA table and confirm
, the coefficient of determination? From the R output identify Multiple R-squared. We find 0.4425. Grab the SSreg and SSTot from the ANOVA table and confirm
![]()
Note 2: The value of ![]() increases with each additional predictor variable included in the model, whether statistically significant contributor or not, regardless of whether the association between the predictor is simply due to chance. The Adjusted R-squared is an attempt to correct for this bias.
increases with each additional predictor variable included in the model, whether statistically significant contributor or not, regardless of whether the association between the predictor is simply due to chance. The Adjusted R-squared is an attempt to correct for this bias.
The residual standard error, RSE, is another model fit statistic.
![]()
The smaller the residual standard error, the better the model fit to the data. See Residual standard error: 1.776 and MSE = 3.1535 above.
And the “statistical significance?” See Pr(>|t|) .There were tests for each coefficient, ![]() and
and ![]() :
:
Table 3. Coefficient estimates.
| P-value | |
| 0.1026 | |
| 0.0255 |
And, at last, note that p-value, Pr(>F), reported in the Analysis of Variance Table, is the same as the p-value for the slope reported as Pr(>|t|).
Make a prediction:
How many matings expected for a 40 g male?
predict(LinearModel.3, data.frame(Body.Mass=40))
R output
1
6.016949
Answer: we predict 6 matings for a 40g male.
See below for predicting confidence limits.
How to extract information from an R object
We can do more. Explore all that is available in the objects LinearModel.3 and myAOV.full with the str() command.
Note 3: str() command lets us look at an object created in R. Type ?str or help(str) to bring up the R documentation. Here, we use str() to look at the structure of the objects we created. “Classes” refers to the R programming class attribute inherited by the object. We first used str() in Chapter 8.3.
str(LinearModel.3)
Output from R
List of 12 $ coefficients : Named num [1:2] -8.475 0.362 ..- attr(*, "names")= chr [1:2] "(Intercept)" "Body.Mass" $ residuals : Named num [1:11] -2.0318 -0.0318 1.9682 0.8814 -1.1186 ... ..- attr(*, "names")= chr [1:11] "1" "2" "3" "4" ... $ effects : Named num [1:11] -12.965 4.746 2.337 1.327 -0.673 ... ..- attr(*, "names")= chr [1:11] "(Intercept)" "Body.Mass" "" "" ... $ rank : int 2 $ fitted.values: Named num [1:11] 2.03 2.03 2.03 3.12 3.12 ... ..- attr(*, "names")= chr [1:11] "1" "2" "3" "4" ... $ assign : int [1:2] 0 1 $ qr :List of 5 ..$ qr : num [1:11, 1:2] -3.317 0.302 0.302 0.302 0.302 ... .. ..- attr(*, "dimnames")=List of 2 .. .. ..$ : chr [1:11] "1" "2" "3" "4" ... .. .. ..$ : chr [1:2] "(Intercept)" "Body.Mass" .. ..- attr(*, "assign")= int [1:2] 0 1 ..$ qraux: num [1:2] 1.3 1.3 ..$ pivot: int [1:2] 1 2 ..$ tol : num 0.0000001 ..$ rank : int 2 ..- attr(*, "class")= chr "qr" $ df.residual : int 9 $ xlevels : Named list() $ call : language lm(formula = Matings ~ Body.Mass, data = bird_matings) $ terms :Classes 'terms', 'formula' language Matings ~ Body.Mass .. ..- attr(*, "variables")= language list(Matings, Body.Mass) .. ..- attr(*, "factors")= int [1:2, 1] 0 1 .. .. ..- attr(*, "dimnames")=List of 2 .. .. .. ..$ : chr [1:2] "Matings" "Body.Mass" .. .. .. ..$ : chr "Body.Mass" .. ..- attr(*, "term.labels")= chr "Body.Mass" .. ..- attr(*, "order")= int 1 .. ..- attr(*, "intercept")= int 1 .. ..- attr(*, "response")= int 1 .. ..- attr(*, ".Environment")=<environment: R_GlobalEnv> .. ..- attr(*, "predvars")= language list(Matings, Body.Mass) .. ..- attr(*, "dataClasses")= Named chr [1:2] "numeric" "numeric" .. .. ..- attr(*, "names")= chr [1:2] "Matings" "Body.Mass" $ model :'data.frame': 11 obs. of 2 variables: ..$ Matings : num [1:11] 0 2 4 4 2 6 3 3 5 8 ... ..$ Body.Mass: num [1:11] 29 29 29 32 32 35 36 38 38 38 ... ..- attr(*, "terms")=Classes 'terms', 'formula' language Matings ~ Body.Mass .. .. ..- attr(*, "variables")= language list(Matings, Body.Mass) .. .. ..- attr(*, "factors")= int [1:2, 1] 0 1 .. .. .. ..- attr(*, "dimnames")=List of 2 .. .. .. .. ..$ : chr [1:2] "Matings" "Body.Mass" .. .. .. .. ..$ : chr "Body.Mass" .. .. ..- attr(*, "term.labels")= chr "Body.Mass" .. .. ..- attr(*, "order")= int 1 .. .. ..- attr(*, "intercept")= int 1 .. .. ..- attr(*, "response")= int 1 .. .. ..- attr(*, ".Environment")=<environment: R_GlobalEnv> .. .. ..- attr(*, "predvars")= language list(Matings, Body.Mass) .. .. ..- attr(*, "dataClasses")= Named chr [1:2] "numeric" "numeric" .. .. .. ..- attr(*, "names")= chr [1:2] "Matings" "Body.Mass" - attr(*, "class")= chr "lm"
Yikes! A lot of the output is simply identifying how R handled our data.frame and applied the lm() function.
Note 4: The “how” refers to the particular algorithms used by R. For example, qraux, pivot, and tol refer to arguments used in the QR decomposition of the matrix (derived from our data.frame) used to solve our linear least squares problem.
However, it’s from this command we learn what is available to extract in our object. For example, what were the actual fitted values (![]() , the predicted Y values) from our linear equation? I marked the relevant
, the predicted Y values) from our linear equation? I marked the relevant str() output above in yellow. We call these from the object with the R code
RegModel.1$fitted.values
and R returns
1 2 3 4 5 6 7 8 9 10 11
2.031780 2.031780 2.031780 3.118644 3.118644 4.205508 4.567797 5.292373 5.292373 5.292373 6.016949
Here’s another look at an object, str(myAOV.full), and how to extract useful values.
Output from R
Classes 'anova' and 'data.frame': 2 obs. of 5 variables:
$ Df : int 1 9
$ Sum Sq : num 22.5 28.4
$ Mean Sq: num 22.53 3.15
$ F value: num 7.14 NA
$ Pr(>F) : num 0.0255 NA
- attr(*, "heading")= chr [1:2] "Analysis of Variance Table\n" "Response: Matings"
Extract the sum of squares: type the object name then $ Sum Sq at the R prompt.
myAOV.full $"Sum Sq"
Output from R
[1] 22.52773 28.38136
Get the residual sum of squares
SSE.myAOV.full <- myAOV.full $"Sum Sq"[2]; SSE.myAOV.full
Output from R
[1] 22.52773
Get the regression sum of squares
SSR.myAOV.full <- myAOV.full $"Sum Sq"[1]; SSR.myAOV.full
Output from R
[1] 50.90909
Now, get the total sums of squares for the model
ssTotal.myAOV.full <- SSE.myAOV.full + SSR.myAOV.full; ssTotal.myAOV.full
Calculate the coefficient of determination
myR_2 <- 1 - (SSE.myAOV.full/(ssTotal.myAOV.full)); myR_2
Output from R
[1] 0.4425091
Which matches what we got before, as it should.
Regression equations may be useful to predict new observations
True. However, you should avoid making estimates beyond the range of the X – values that were used to calculate the best fit regression equation! Why? The answer has to do with the shape of the confidence interval around the regression line.
I’ve drawn an exaggerated confidence interval (CI), for a regression line between an X and a Y variable. Note that the CI is narrow in the middle, but wider at the end. Thus, we have more confidence in predicting new Y values for data that fall within the original data because this is the region where we are most confident.
Calculating the CI for the linear model follows from CI calculations for other estimates. It is a simple concept — both the intercept and slope were estimated with error, so we combine these into a way to generalize our confidence in the regression model as a whole given the error in slope and intercept estimation.
![]()
The calculation of confidence interval for the linear regression involves the standard error of the residuals, the sample size, and expressions relating the standard deviation of the predictor variable X — we use the t distribution.
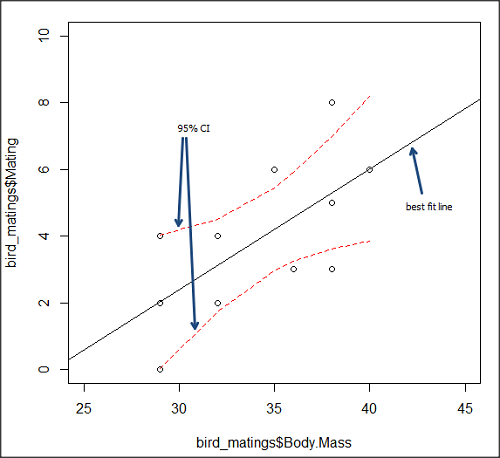
Figure 5. 95% confidence interval about the best fit line.
How I got this graph
plot(bird_matings$Body.Mass,bird_matings$Mating,xlim=c(25,45),ylim=c(0,10))
mylm <- lm(bird_matings$Mating~bird_matings$Body.Mass)
predict(mylm, interval = c("confidence"))
abline(mylm, col = "black")
x<-bird_matings$Body.Mass
lines(x, prd[,2], col= "red", lty=2)
lines(x, prd[,3], col= "red", lty=2)
Nothing wrong with my code, but getting all of this to work in R might best be accomplished by adding another package, a plug-in for Rcmdr called RcmdrPlugin.HH. HH refers to “Heiberger and Holland,” and provides software support for their textbook Statistical Analysis and Data Display, 2nd edition.
Assumptions of OLS, an introduction
We will cover assumptions of Ordinary Least Squares in detail in 17.8, Assumptions and model diagnostics for Simple Linear Regression. For now, briefly, the assumptions for OLS regression include:
- Linear model is appropriate: the data are well described (fit) by a linear model
- Independent values of Y and equal variances. Although there can be more than one Y for any value of X, the Y’s cannot be related to each other (that’s what we mean by independent). Since we allow for multiple Y’s for each X, then we assume that the variances of the range of Ys are equal for each X value (this is similar to our ANOVA assumptions for equal variance by groups).
- Normality. For each X value there is a normal distribution of Ys (think of doing the experiment over and over)
- Error (residuals) are normally distributed with a mean of zero.
Additionally, we assume that measurement of X is done without error (the equivalent, but less restrictive practical application of this assumption is that the error in X is at least negligible compared to the measurements in the dependent variable). Multiple regression makes one more assumption, about the relationship between the predictor variables (the X variables). The assumption is that there is no multicolinearity: the X variables are not related or associated to each other.
In some sense the first assumption is obvious if not trivial — of course a “line” needs to fit the data so why not plow ahead with the OLS regression method, which has desirable statistical properties and let the estimation of slopes, intercept and fit statistics guide us? One of the really good things about statistics is that you can readily test your intuition about a particular method using data simulated to meet, or not to meet, assumptions.
What about assumptions and BLUE? Interestingly, OLS coefficient estimates remain unbiased even if the errors are not normal, as long as other assumptions (like a mean of zero, no correlation, and constant variance) hold. However, the OLS estimators are no longer the “best” linear unbiased estimators (BLUE) because the efficiency property is lost. As we have discussed before, the main consequence is the effects on statistical inference. Without normality, standard errors, t-statistics, confidence intervals, and p-values will be unreliable.
Note 5: The efficiency property means the estimator has the smallest variance among all linear, unbiased estimators. And “unbiased,” again, means the estimator’s expected value is equal to the true parameter value.
Coming up with datasets like these can be tricky for beginners. Thankfully, others have stepped in and provide tools useful for data simulations which greatly facilitate the kinds of testing of assumptions — and more importantly, how assumptions influence our conclusions — statisticians greatly encourage us all to do (see Chatterjee and Firat 2007).
Questions
- Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
- Predict new values for number of matings for male birds of 20g and 60g.
- Anscombe’s quartet (Anscombe 1973) is a famous example of this approach and the fictitious data can be used to illustrate the fallacy of relying solely on fit statistics and coefficient estimates.
Here are the data (modified from Anscombe 1973, p. 19) — I leave it to you to discover the message by using linear regression on Anscombe’s data set. Hint: play naïve and generate the appropriate descriptive statistics, make scatterplots for each X, Y set, then run regression statistics, first on each of the X, Y pairs (there were four sets of X, Y pairs).
Anscombe’s quartet
| X | Y1 | Y2 | Y3 | Y4 |
|---|---|---|---|---|
| 10 | 8.04 | 9.14 | 7.46 | 6.58 |
| 8 | 6.95 | 8.14 | 6.77 | 5.76 |
| 13 | 7.58 | 8.74 | 12.74 | 7.71 |
| 9 | 8.81 | 8.77 | 7.11 | 8.84 |
| 11 | 8.33 | 9.26 | 7.81 | 8.47 |
| 14 | 9.96 | 8.10 | 8.84 | 7.04 |
| 6 | 7.24 | 6.13 | 6.08 | 5.25 |
| 4 | 4.26 | 3.10 | 5.39 | 12.50 |
| 12 | 10.84 | 9.13 | 8.15 | 5.56 |
| 7 | 4.82 | 7.26 | 6.42 | 7.91 |
| 5 | 5.68 | 4.74 | 5.73 | 6.89 |
| Mean (±SD) | 7.50 (2.032) | 7.50 (2.032) | 7.50 (2.032) | 7.50 (2.032) |
Quiz Chapter 17.1
Ordinary Least Squares
Chapter 17 contents
- Introduction
- Simple Linear Regression
- Relationship between the slope and the correlation
- Estimation of linear regression coefficients
- OLS, RMA, and smoothing functions
- Testing regression coefficients
- ANCOVA – analysis of covariance
- Regression model fit
- Assumptions and model diagnostics for Simple Linear Regression
- References and suggested readings (Ch17 & 18)
16.6 – Similarity and Distance
Introduction
A measure of dependence between two random variables. Unlike Pearson Product Moment correlation, distance correlation measures strength of association between the variables whether or not the relationship is linear. Distance correlation is a recent addition to the literature, first reported by Gábor J. Székely (e.g., Székely et al. 2007). The package correlation (Makowski et al 2019) offers distance correlation and significance test.
Example, fly wing dataset introduced 16.1 – Product moment correlation
library(correlation) Area <- c(0.446, 0.876, 0.390, 0.510, 0.736, 0.453, 0.882, 0.394, 0.503, 0.535, 0.441, 0.889, 0.391, 0.514, 0.546, 0.453, 0.882, 0.394, 0.503, 0.535) Length <- c(1.524, 2.202, 1.520, 1.620, 1.710, 1.551, 2.228, 1.460, 1.659, 1.719, 1.534, 2.223, 1.490, 1.633, 1.704, 1.551, 2.228, 1.468, 1.659, 1.719) FlyWings <- data.frame(Area, Length) correlation(FlyWings,method="distance")
Output from R
# Correlation Matrix (distance-method)
Parameter1 | Parameter2 | r | 95% CI | t(169) | p
------------------------------------------------------------------
Area | Length | 0.92 | [0.80, 0.97] | 30.47 | < .001***
p-value adjustment method: Holm (1979)
Observations: 20
The product-moment correlation was 0.97 with 95% confidence interval (0.92, 0.99). The note about “p-value adjustment method: Holm (1979)” refers to the algorithm used to mitigate the multicomparison problem, which we first introduced in Chapter 12.1. The correction is necessary in this context because of how the algorithm conducts the test of the distance correlation. Please see Székely and Rizzo (2013) for more details.
Which should you report? For cases where it makes sense to test for a linear association, then the Product-moment correlation is the one to use. For other cases where no inference of linearity is expected, then the distance correlation makes sense.
Similarity and Distance
Similarity and distance are related mathematically. When two things are similar, the distance between them is small; When two things are dissimilar, the distance between them is great. Whether similarity (sometimes dissimilarity) or distance, the estimate is a statistic. The difference between the two is that the typical distance measures one sees in biostatistics all obey the triangle inequality rule while similarity (dissimilarity) indices do not necessarily obey the triangle inequality rule.
Distance measures
Distance is a way to talk about how far (or how close) two objects are from each other (Fig. 1). The distance may be relate to physical distance (map distance), or in mathematics, distance is a metric or statistic. Euclidean distance is the distance between two points in either the X-Y plane or 3-dimensional space measures the length of a segment connecting the two points (e.g., x1,y1 = 1, 4 and x2,y2 = 1,4).
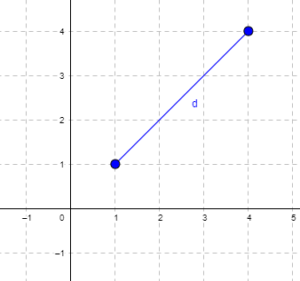
Figure 1. Cartesian plot of two points, the first at x1 = 1 and y1 = 1 and the second at x2 = 4 and y2 = 4.
For two points (x1, y1 and x2, y2) described in two dimensions (e.g., an xy plane), the distance d is given by
![]()
For two points described in three (e.g., an xyz-space), or more dimensions, the distance d is given by
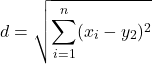
Distances of this form are Euclidean distances and can be directly obtained by use of the Pythagorean Theorem. The triangle inequality rule then applies (i.e., the sum of any two sides must be less than the length of the remaining side). Euclidean distance measures also include
- Manhattan distance: the sum of absolute difference between the measures in all dimensions of two points.
![]()
- Chebyshev distance: also called the maximum value distance, the distance between two points is the greatest of their differences along any coordinate dimension.
![]()
Note: We first met Chebyshev in Chapter 3.5.
Example
There are a number of distance measures. Let’s begin discussion of distance with geographic distance as an example. Consider the distances between cities (Table 1).
Table 1. Distances (miles) among cities.
| Honolulu | Seattle | Manila | Tokyo | Houston | |
| Honolulu | 0 | 2667.57 | 5323.37 | 3849.99 | 3891.82 |
| Seattle | 2667.57 | 0 | 6590.23 | 4776.81 | 1888.06 |
| Manilla | 5323.37 | 6590.23 | 0 | 1835.1 | 8471.48 |
| Tokyo | 3849.99 | 4776.81 | 1835.1 | 0 | 6664.82 |
| Houston | 3891.82 | 1888.06 | 8471.48 | 6664.82 | 0 |
This table is a distance matrix — note that along the diagonal are “zeros,” which should make sense — the distance between an object and itself is, well zero. Above and below the diagonal you see the distance between one city and another. This is a special kind of matrix called a symmetric matrix. Enter the distance in miles (1 mile = 1.609344) between 2 cities (this is “pairwise“). There are many resources “out there,” to help you with this. For example, I found a web site called mapcrow that allowed me to enter the cities and calculate distances between them.
To get the distance matrix, use this online resource, the Geographic Distance Matrix Calculator.
Real world problem, use geodist package. Provide latitude and longitude coordinates
The dist() function in R base will return Manhattan distance and others () on matrices.
R script
HNL <- c(0, 2667.57, 5323.37, 3849.99, 3891.82)
SEA <- c(2667.57, 0, 6590.23, 4776.81, 1888.06)
MAN <- c(5323.37, 6590.23, 0, 1835.1, 8471.48)
TOK <- c(3849.99, 4776.81, 1835.1, 0, 6664.82)
HOU <- c(3891.82, 1888.06, 8471.48, 6664.82, 0)
mat.city <- rbind(HNL, SEA, MAN, TOK, HOU)
colnames(mat.city) <- c("HNL", "SEA", "MAN", "TOK", "HOU")
mat.city
dist(mat.city, method = "manhattan")
R output
HNL SEA MAN TOK SEA 9532.58 MAN 21163.95 25361.39 TOK 16070.49 20267.93 8763.66 HOU 14526.09 8769.63 27906.40 22896.60
Other distances, replace “manhattan” with
For example,
dist(mat.city, method = "euclidean")
R output
HNL SEA MAN TOK SEA 4550.917 MAN 9853.774 12079.347 TOK 7345.155 9615.750 3931.736 HOU 6980.976 3966.360 13820.922 11883.931
The Chebyshev distance doesn’t seem to be part of base R, but is available in other packages (philentropy, function distance).
Note 1. Need a different distance measure? Chances are philentropy packages can calculate it; use getDistMethods() to explore available measures.
R script
require(philentropy) distance(mat.city, method = "chebyshev", diag=FALSE, upper=FALSE)
R output
v1 v2 v3 v4 v5 v1 0.00 2667.57 5323.37 3849.99 3891.82 v2 2667.57 0.00 6590.23 4776.81 1888.06 v3 5323.37 6590.23 0.00 1835.10 8471.48 v4 3849.99 4776.81 1835.10 0.00 6664.82 v5 3891.82 1888.06 8471.48 6664.82 0.00
Distance measures used in biology
It is easy to see how the concept of genetic distance between a group of species (or populations within a species) could be used to help build a network, with genetically similar species grouped together and genetically distant species represented in a way to represent how far removed they are from each other. Here, we speak of distance as in similarity: two species (populations) that are similar are close together, and the distance between them is short. In contrast, two species (populations) that are not similar would be represented by a great distance between them. Genetic distance is the amount of divergence of species from each other. Smaller genetic distances reflects close genetic relationship. Hamming distance is a common choice, good for categorical data.
Here’s an example (Fig. 2), RAPD gel for five kinds of beans. RAPD stands for random amplified polymorphic DNA.
Figure 2. RAPD gel (simulated) five kinds of beans.
Samples were small red beans (SRB), garbanzo beans (GB), split green pea (SGP), baby lima beans (BLB), and black eye peas (BEP). RAPD primer 1 was applied to samples in lanes 1 – 6; RAPD primer 2 was applied to samples in lane 7 – 12. Lane 1 & 7 = SRB; Lane 2 & 8 = GB; Lanes 3 & 9 = SGP; Lane 4 & 10 = BLB; Lane 5 & 11 = BB; Lane 6 & 12 = BEP.
Here’s how to go from gel documentation to the information needed for genetic distance calculations (see below). I’ll use “1” to indicate presence of a band, “0” to indicate absence of a band, and “?” to indicate no information. For simplicity, I ignored the RF value, but ranked the bands by order of largest (= 1) to smallest (=8) fragment.
We need three pieces of information from the gel to calculate genetic distance.
NA = the number of markers for taxon A
NB = the number of markers for taxon B
NAB = the number of markers in common between A and B (this is the pairwise part — we are comparing taxa two at a time.
First, compare the beans against the same primer. My results for primer 1 are in Table 2; results for primer 2 are in Table 3.
Table 2. Bands for Primer 1
| marker | lane 1 | Lane 2 | Lane3 | Lane 4 | Lane 5 | Lane 6 |
| 1 | 1 | 1 | ? | 1 | 0 | 0 |
| 3 | 1 | 1 | ? | 0 | 0 | 0 |
| 5 | 1 | 1 | ? | 1 | 1 | 1 |
| 7 | 0 | 0 | ? | 0 | 1 | 1 |
Table 3 for Primer 2
| marker | Lane 7 | Lane 8 | Lane 9 | Lane 10 | Lane 11 | Lane 12 |
| 2 | 0 | 1 | ? | 0 | 0 | 0 |
| 4 | 1 | 0 | ? | 0 | 1 | 0 |
| 6 | 0 | 1 | ? | 0 | 1 | 0 |
| 8 | 1 | 0 | ? | 1 | 1 | 1 |
From Table 2 and Table 3, count NA (= NB ) for each taxon. Results are in Table 4.
Table 4. Bands for each taxon.
| Taxon | No. markers from Primer1 |
No. markers from Primer2 |
Total |
| SRB | 3 | 2 | 5 |
| GB | 3 | 2 | 5 |
| SGP | ? | ? | ? |
| BLB | 2 | 1 | 3 |
| BB | 2 | 3 | 5 |
| BEP | 2 | 1 | 3 |
To get Hamming distance between SRB and BB (from Table 2 and Table 3), R script as follows.
SRB <- c(1, 1, 1, 0, 0, 1, 0, 1) GB <- c(0, 0, 1, 1, 0, 1, 1, 1) # sum the inequality between the two vectors sum(SRB != GB)
R output
> sum(SRB != GB) [1] 4
Hamming distance between small red beans and black beans for the markers is 4.
Note 2. h/t https://www.r-bloggers.com/2021/12/how-to-calculate-hamming-distance-in-r/
As you can see, there is no simple relationship among the taxa; there is no obvious combination of markers that readily group the taxa by similarity. So, I need a computer to help me. I need a measure of genetic distance, a measure that indicates how (dis)similar the different varieties are for our genetic markers. I’ll use a distance calculation that counts only the “present” markers, not the “absent” markers, which is more appropriate for RAPD. I need to get the NAB values, the number of shared markers between pairs of taxa.
Table 5. NAB
| SRB | GB | BLB | BB | BEP | |
| SRB | 0 | 3 | 3 | 3 | 2 |
| GB | 0 | 2 | 2 | 1 | |
| BLB | 0 | 2 | 2 | ||
| BB | 0 | 3 | |||
| BEP | 0 |
The equation for calculating Nei’s distance is:
![]()
where NA = number of bands in taxon “A”, NB = number of bands in taxon “B”, and NAB is the number of bands in common between A and B (Nei and Li 1979). Here’s an example calculation.
Let A = SRB and B = GB, then
![]()
The R package adegenet includes Nei’s distance (and others), but the data set assumes more information than I provided here (e.g., ploidy, alleles).
Which distance measure to use?
Unfortunately, not a simple answer. A quick search of journals will likely return “Euclidean distance” the most common (Evolution = 328, Ecology = 298). The different distance (similarity) measures were developed to solve sometimes related, but often different problems, and so a review of the statistical properties would be warranted before making a choice. Helpful reviews
Questions
1. Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
2. Review all of the different correlation estimates we introduced in Chapter 16 and construct a table to help you learn. Product moment correlation is presented as example.
| Name of correlation | variable 1 type | variable 2 type | purpose |
| Product moment | ratio | ratio | estimate linear association |
3. Be able to define and distinguish between similarity measures, dissimilarity measures quantify, and distance.
4. Return to our counts of colored candies from mini bags of Skittles, M&M plain chocolate, and M&M peanut. Which two are most similar? Here’s a data sample. Calculate the Euclidean pairwise distances, then the Manhattan distances among the bags of candies. Do you get the same answer?
The data
| Color | Skittles | mm_plain | mm_peanut |
|---|---|---|---|
| Blue | 0 | 19 | 18 |
| Green | 40 | 43 | 28 |
| Yellow | 30 | 29 | 9 |
| Orange | 26 | 39 | 18 |
| Red | 28 | 15 | 4 |
| Purple | 28 | 0 | 4 |
| Brown | 0 | 16 | 0 |
Example R code
Note 3. The code works. Remember, sometimes when copying code from a website you pickup additional characters. While every effort has been made to provide clean html — if you run into problems, chances are the simplest fix is to type the code fresh rather than trying to figure out why the copied code fails.
mySites <- data.frame(
Site <- c("A","B","C"),
Spp1 <- c(10, 5, 12),
Spp2 <- c(7, 15, 8),
Spp3 <- c(3, 2, 9)
)
# View the matrix print(mySites)
# Exclude the "Site" column myAbundances = mySites[,-1]
# Calculate the Euclidean distance matrix dist(myAbundances,method="euclidean")
The output should be
1 2 2 9.486833 3 6.403124 12.124356
where “1” refers to site A, “2” refers to site B, and “3” refers to site C. We would conclude that sites A and C are most similar, or have similar species abundance, because they have the smallest Euclidean distance. As you can imagine, there’s more to interpreting these statistics, and in particular, Euclidean distance can lead to some confusing interpretations about abundance and species composition between sites, eg, Orlóci paradox. Interested readers should see Ricotta 2021.
Note 4. Orlóci paradox refers to a problem with Euclidean distances: two sites with no species in common may be interpreted as more similar that two sites that share the same species. This is because Euclidean distance is more sensitive to abundance than presence/absence.
Why then teach Euclidean distance? The Euclidean distance is a standard measure, it helps lay the foundation for understanding geometric concepts. Its limitations are a reminder that students need to be aware that alternative methods are sometimes needed.
Quiz Chapter 16.6
Similarity and Distance
Chapter 16 contents
16.4 – Spearman and other correlations
Introduction
Pearson product moment correlation is used to describe the level of linear association between two variables. There are many types of correlation estimators in addition to the familiar Product Moment Correlation, r.
Spearman rank correlation
If you take the ranks for X 1 and the ranks for X 2, the correlation of ranks is called Spearman rank correlation, rs. Spearman correlation is a nonparametric statistic. Like the product moment correlation, it can take values between -1 and +1.
For variables X 1 and X 2, the rank order correlation may be calculated on the ranks as
![]()
where di is the difference between the ranks of X 1 and X 2 for each experimental unit. This formula assumes that there are no tied ranks; if there are, use the equation for the product moment correlation instead (but on the ranks).
R commander has an option to calculate the Spearman rank correlation simply by selecting the check box in the correlation sub menu. However, if the data set is small, it may be easier to just run the correlation in the script window.
Our example for the product moment correlation was between Drosophila fly wing length and wing area (Table 1).
Table 1. Fly wing lengths and area, units mm and mm2, respectively (Dohm pers obs.)
| Obs | Student | Length | Area |
|---|---|---|---|
| 1 | S01 | 1.524 | 0.446 |
| 2 | S01 | 2.202 | 0.876 |
| 3 | S01 | 1.52 | 0.39 |
| 4 | S01 | 1.62 | 0.51 |
| 5 | S01 | 1.71 | 0.736 |
| 6 | S03 | 1.551 | 0.453 |
| 7 | S03 | 2.228 | 0.882 |
| 8 | S03 | 1.46 | 0.394 |
| 9 | S03 | 1.659 | 0.503 |
| 10 | S03 | 1.719 | 0.535 |
| 11 | S05 | 1.534 | 0.441 |
| 12 | S05 | 2.223 | 0.889 |
| 13 | S05 | 1.49 | 0.391 |
| 14 | S05 | 1.633 | 0.514 |
| 15 | S05 | 1.704 | 0.546 |
| 16 | S08 | 1.551 | 0.453 |
| 17 | S08 | 2.228 | 0.882 |
| 18 | S08 | 1.468 | 0.394 |
| 19 | S08 | 1.659 | 0.503 |
| 20 | S08 | 1.719 | 0.535 |
Repeated observations by image analysis (ImageJ) were collected by four students (S01, S03, So5, S08) from fixed wings to glass slides.
Here’s the scatterplot of the ranks of fly wing length and fly wing area (Fig 1).
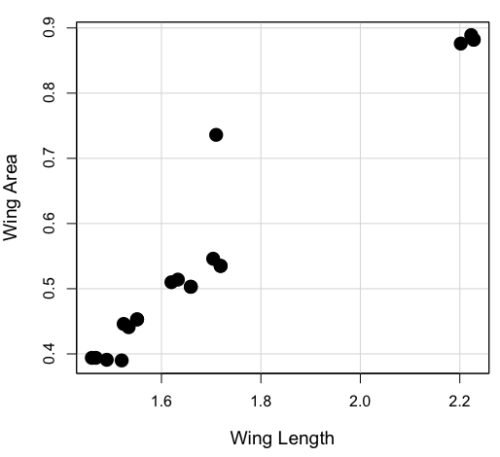
Figure 1. Drosophila wing area (mm2) by wing length (mm).
A nonparametric alternative to the product moment correlation, the Spearman Rank correlation can be obtained directly. The Spearman correlation involves ranking the data, ie, converting data types, from ratio scale data to ordinal scale, then applying the same formula used for the Product moment correlation to the ranked data. The Spearman correlation would be the choice for testing linear association between two ordinal type variables. It is also appropriate in lieu of the parametric product moment correlation when the statistical assumptions are not met, eg, normality assumption.
R code
For the Spearman rank correlation, at the R prompt type
cor.test(Area, Length, alternative="two.sided", method="spearman") R returns with Spearman's rank correlation rho data: Area and Length S = 58.699, p-value = 5.139e-11 alternative hypothesis: true rho is not equal to 0 sample estimates: rho 0.9558658
Alternatively, to calculate either correlation, use R Commander.
Rcmdr: Statistics → Summaries→ Correlation test
Example
BM=c(29,29,29,32,32,35,36,38,38,38,40) Matings=c(0,2,4,4,2,6,3,3,5,8,6) cor.test(BM,Matings, method="spearman") Warning in cor.test.default(BM, Matings, method = "spearman") : Cannot compute exact p-value with ties Spearman's rank correlation rho data: BM and Matings S = 77.7888, p-value = 0.03163 alternative hypothesis: true rho is not equal to 0 sample estimates: rho 0.6464143
cor.test(BM,Matings, method="pearson") Pearson's product-moment correlation data: BM and Matings t = 2.6728, df = 9, p-value = 0.02551 alternative hypothesis: true correlation is not equal to 0 95 percent confidence interval: 0.1087245 0.9042515 sample estimates: cor 0.6652136
Other correlations
Kendall’s tau
Another nonparametric correlation is Kendall’s tau (τ). Rank the X 1 values, then rank the X 2 values. Count the number of X 1, X 2 pairs that have the same rank (concordant pairs) and the number of X 1, X 2 pairs that do not have the same rank (discordant pairs), Kendall’s tau is then
![]()
where n is the number of pairs.
Note 1: The denominator is our familiar number of pairwise comparisons if we take k = n
![]()
We introduced concordant and discordant pairs when we presented McNemar’s test and cross-classified experimental design in Chapter 9.6.
Example: Judging of Science Fair posters
What is the agreement between two judges, A and B, who evaluated the same science fair posters? Posters were evaluated on if the student’s project was hypothesis-based and judges used a Likert-like scale Strongly disagree (1), Somewhat disagree (2), Neutral (3), Somewhat agree (4), Strongly agree (5).
Table 2. Two judges evaluated six posters for evidence of hypothesis-based project
| Poster | Judge.A | Judge.B |
| 1 | 5 | 4 |
| 2 | 2 | 3 |
| 3 | 4 | 2 |
| 4 | 3 | 1 |
| 5 | 2 | 1 |
| 6 | 4 | 3 |
A concordant pair represents a poster ranked higher by both judges, while a discordant pair is a poster ranked high by one judge but low by another judge. Poster 1 and poster 5 were concordant pairs,
In R, it is simple to get this correlation directly by invoking the cor.test function and specifying the method equal to kendall. The cor.test assumes that the data are in a matrix, so use the cbind function to bind two vectors together – note the vectors need to have the same number of observations. If the data set is small, it is easier to just enter the data directly in the script window of R commander.
A = c(2,2,3,4,4,5) B = c(1,3,1,2,3,4) m = cbind(A,B) cor.test(A,B, method="kendall") Cannot compute exact p-value with ties Kendall's rank correlation tau data: A and B z = 1.4113, p-value = 0.1581 alternative hypothesis: true tau is not equal to 0 sample estimates: tau 0.5384615
End of R output
There were no ties in this data set, but we can run the product moment correlation just for comparison
cor.test(A,B, method="pearson") Pearson's product-moment correlation data: A and B t = 1.4649, df = 4, p-value = 0.2168 alternative hypothesis: true correlation is not equal to 0 95 percent confidence interval: -0.4239715 0.9478976 sample estimates: cor 0.5909091
End R output
Tetrachoric and Polychoric correlation
Tetrachoric correlations used for binomial outcomes (yes, no), polychoric correlation used for ordinal categorical data like the Likert scale. Introduced by Karl Pearson, commonly applied correlation estimate for Item Analysis in psychometric research. Pyschometrics, a sub-discipline within psychology and now a significant part of education research, is about evaluating assessment tools.
R package psych.
R code: Tetrachoric correlation
tetrachoric(x,y=NULL,correct=.5,smooth=TRUE,global=TRUE,weight=NULL,na.rm=TRUE,
delete=TRUE)
R code: Polychoric correlation
polychoric(x,smooth=TRUE,global=TRUE,polycor=FALSE,ML=FALSE, std.err=FALSE,
weight=NULL,correct=.5,progress=TRUE,na.rm=TRUE, delete=TRUE)
Polyserial correlation
R package polychor. Used to estimate linear association between a ratio scale variable and an ordinal variable.
R code: Polyserial correlation
polyserial(x,y)
Biserial correlation would be a special case of the polyserial correlation, where ordinal variable is replaced by a dichotomous (binomial) variable.
R code: Polyserial correlation
biserial(x,y)
Intra-class correlation coefficient
Both the ICC and the product moment correlation, r, which we introduced in Chapter 16.1, are measures of strength of linear association between two ratio scale variables (Jinyuan et al 2016). But ICC is more appropriate for association between repeat measures of the same thing, eg, repeat measures of running speed. In contrast, the product moment correlation can be used to describe association between any two variables, eg, between repeat measures of running speed, but also between say running speed and maximum jumping height. ICC is used when quantitative measures are organized into paired groups, eg, before and after on same subjects, or cross-classified designs. ICC was introduced in Chapter 12.3 as part of discussion of repeated measures and random effects. ICC is used extensively to assess reliability of a measurement instrument (Shrout and Fleiss 1979; McGraw and Wong 1996).
Example. Data from Table 2
library(psych)
ICC(myJudge, lmer=FALSE)
R output follows
Intraclass correlation coefficients
type ICC F df1 df2 p lower bound upper bound
Single_raters_absolute ICC1 0.40 2.3 5 6 0.166 -0.306 0.84
Single_random_raters ICC2 0.46 3.9 5 5 0.081 -0.093 0.85
Single_fixed_raters ICC3 0.59 3.9 5 5 0.081 -0.130 0.90
Average_raters_absolute ICC1k 0.57 2.3 5 6 0.166 -0.880 0.91
Average_random_raters ICC2k 0.63 3.9 5 5 0.081 -0.205 0.92
Average_fixed_raters ICC3k 0.74 3.9 5 5 0.081 -0.299 0.95
Number of subjects = 6 Number of Judges = 2
See the help file for a discussion of the other 4 McGraw and Wong estimates
Lots of output, lots of “ICC”. However, rather than explaining each entry, reflect on the type and review the data. Were the posters evaluated repeatedly? Posters were evaluated twice, but only once per judge, so there is a repeated design with respect to the posters. Had the judges been randomly selected from a population of all possible judges? No evidence was provided to suggest this, so judges were a fixed factor (see Chapter 12.3 and Chapter 14.3).
The six ICC estimates reported by R follow discussion in Shrout and Fliess (1979), and our description fits their Case 3: “Each target is rated by each of the same k judges, who are the only judges of interest (p. 421)” Thus, we find ICC for single fixed rater, ICC = 0.59. Note that we would fail to reject the hypothesis that the judges evaluations were associated.
Questions
1. Write up three learning outcomes for this page. Hint: Point your favorite generative AI to this page and ask for help.
2. See Homework 10, Mike’s Workbook for biostatistics
Quiz Chapter 16.4
Spearman and other correlations